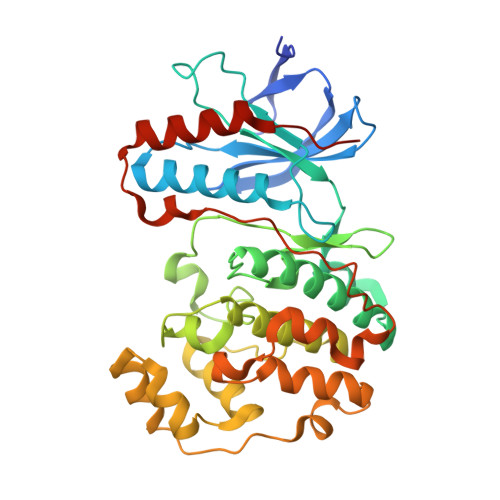
Structural basis for p38alpha MAP kinase quinazolinone and pyridol-pyrimidine inhibitor specificity
Fitzgerald, C.E., Patel, S.B., Becker, J.W., Cameron, P.M., Zaller, D., Pikounis, V.B., O'Keefe, S.J., Scapin, G.(2003) Nat Struct Biol 10: 764-769
- PubMed: 12897767 Search on PubMed
- DOI: https://doi.org/10.1038/nsb949
- Primary Citation Related Structures:
1OUK, 1OUY, 1OVE - PubMed Abstract:
The quinazolinone and pyridol-pyrimidine classes of p38 MAP kinase inhibitors have a previously unseen degree of specificity for p38 over other MAP kinases. Comparison of the crystal structures of p38 bound to four different compounds shows that binding of the more specific molecules is characterized by a peptide flip between Met109 and Gly110. Gly110 is a residue specific to the alpha, beta and gamma isoforms of p38. The delta isoform and the other MAP kinases have bulkier residues in this position. These residues would likely make the peptide flip energetically unfavorable, thus explaining the selectivity of binding. To test this hypothesis, we constructed G110A and G110D mutants of p38 and measured the potency of several compounds against them. The results confirm that the selectivity of quinazolinones and pyridol-pyrimidines results from the presence of a glycine in position 110. This unique mode of binding may be exploited in the design of new p38 inhibitors.
- Department of Immunology, Merck Research Laboratories, PO Box 2000, Rahway, New Jersey 07065, USA.
Organizational Affiliation: