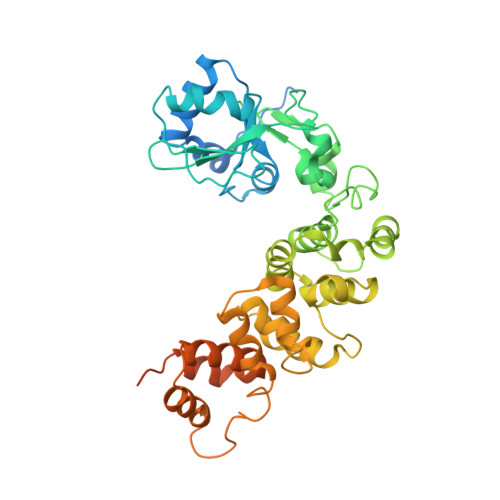
Crystal Structure of the Human CCA-adding Enzyme: Insights into Template-independent Polymerization
Augustin, M.A., Reichert, A.S., Betat, H., Huber, R., Moerl, M., Steegborn, C.(2003) J Mol Biology 328: 985-994
- PubMed: 12729736
- DOI: https://doi.org/10.1016/s0022-2836(03)00381-4
- Primary Citation Related Structures:
1OU5 - PubMed Abstract:
All tRNA molecules carry the invariant sequence CCA at their 3'-terminus for amino acid attachment. The post-transcriptional addition of CCA is carried out by ATP(CTP):tRNA nucleotidyltransferase, also called CCase. This enzyme catalyses a unique template-independent but sequence-specific nucleotide polymerization reaction. In order to reveal the molecular mechanism of this activity, we solved the crystal structure of human CCase by single isomorphous replacement. The structure reveals a four domain architecture with a cluster of conserved residues forming a positively charged cleft between the first two domains. Structural homology of the N-terminal CCase domain to other nucleotidyltransferases could be exploited for modeling a tRNA-substrate complex. The model places the tRNA 3'-end into the N-terminal nucleotidyltransferase site, close to a patch of conserved residues that provide the binding sites for CTP and ATP. Based on our results, we introduce a corkscrew model for CCA addition that includes a fixed active site and a traveling tRNA-binding region formed by flexible parts of the protein.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18A, D-82152 Martinsried, Germany. augustin@biochem.mpg.de
Organizational Affiliation: