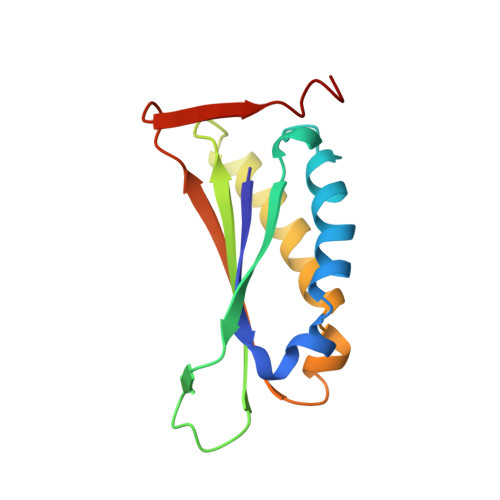
Enzymatic ketonization of 2-hydroxymuconate: specificity and mechanism investigated by the crystal structures of two isomerases.
Subramanya, H.S., Roper, D.I., Dauter, Z., Dodson, E.J., Davies, G.J., Wilson, K.S., Wigley, D.B.(1996) Biochemistry 35: 792-802
- PubMed: 8547259 Search on PubMed
- DOI: https://doi.org/10.1021/bi951732k
- Primary Citation Related Structures:
1OTF, 1OTG - PubMed Abstract:
5-Carboxymethyl-2-hydroxymuconate isomerase (CHMI) and 4-oxalocrotonate tautomerase (4-OT) are enzymes that catalyze the isomerization of unsaturated ketones. They share a common enzyme mechanism, although they show a preference for different substrates. There is no apparent sequence homology between the enzymes. To investigate the molecular mechanism and the basis for their substrate specificity, we have determined the crystal structures of the two enzymes at high resolution. 4-OT is hexameric, with the subunits arranged with 32 symmetry. CHMI is trimeric and has extensive contacts between subunits, which include secondary structural elements. The central core of the CHMI monomer has a fold similar to a 4-OT dimer, but the secondary structural elements that form the subunit contacts around the 3-fold axis are different in the two enzymes. The region of greatest similarity between the two enzymes is a large pocket that is proposed to be the active site. The enzymes appear to operate via a "one-base" mechanism, and the possible role of residues in this pocket is discussed in view of this idea. Finally, the molecular basis for substrate specificity in the two enzymes is discussed.
- Laboratory of Molecular Biophysics, University of Oxford, U.K.
Organizational Affiliation: