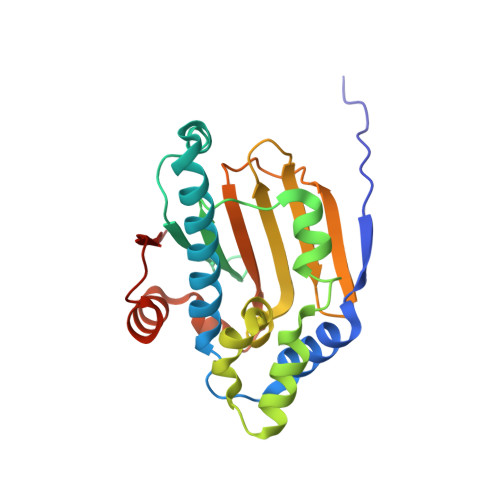
Crystal Structure and Molecular Modeling of 17-DMAG in Complex with Human Hsp90
Jez, J.M., Chen, J.C., Rastelli, G., Stroud, R.M., Santi, D.V.(2003) Chem Biol 10: 361-368
- PubMed: 12725864 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-5521(03)00075-9
- Primary Citation Related Structures:
1OSF - PubMed Abstract:
Hsp90 is an attractive chemotherapeutic target because it chaperones the folding of proteins found in multiple signal transduction pathways. We describe the 1.75 A resolution crystal structure of human Hsp90 alpha (residues 9-236) complexed with 17-desmethoxy-17-N,N-dimethylaminoethylamino-geldanamycin (17-DMAG). The structure revealed an altered set of interactions between the 17-substituent and the protein compared to geldanamycin and the 17-dimethylaminoethyl moiety pointing into solvent, but otherwise was similar to that reported for the complex with geldanamycin. Targeted molecular dynamics simulations and energetic analysis indicate that geldanamycin undergoes two major conformational changes when it binds Hsp90, with the key step of the conversion being the trans to cis conformational change of the macrocycle amide bond. We speculate that 17-DMAG analogs constrained to a cis-amide in the ground state could provide a significant increase in affinity for Hsp90.
- Kosan Biosciences, Inc., 3832 Bay Center Place, Hayward, CA 94545, USA.
Organizational Affiliation: