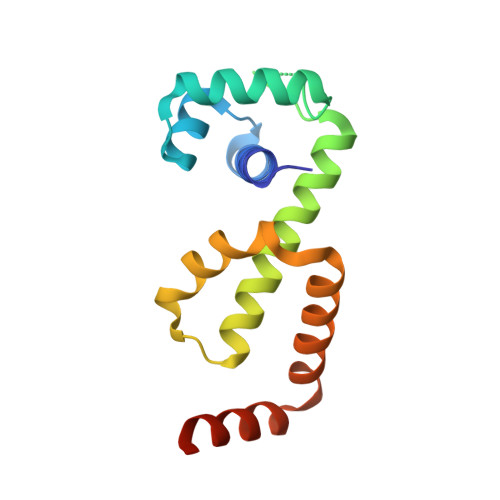
Structure of the Manganese-Bound Manganese Transport Regulator of Bacillus subtilis
Glasfeld, A., Guedon, E., Helmann, J.D., Brennan, R.G.(2003) Nat Struct Biol 10: 652-657
- PubMed: 12847518 Search on PubMed
- DOI: https://doi.org/10.1038/nsb951
- Primary Citation Related Structures:
1ON1, 1ON2 - PubMed Abstract:
The Bacillus subtilis manganese transport regulator, MntR, binds Mn2+ as an effector and is a repressor of transporters that import manganese. A member of the diphtheria toxin repressor (DtxR) family of metalloregulatory proteins, MntR exhibits selectivity for Mn2+ over Fe2+. Replacement of a metal-binding residue, Asp8, with methionine (D8M) relaxes this specificity. We report here the X-ray crystal structures of wild-type MntR and the D8M mutant bound to manganese with 1.75 A and 1.61 A resolution, respectively. The 142-residue MntR homodimer has substantial structural similarity to the 226-residue DtxR but lacks the C-terminal SH3-like domain of DtxR. The metal-binding pockets of MntR and DtxR are substantially different. The cation-to-cation distance between the two manganese ions bound by MntR is 3.3 A, whereas that between the metal ions bound by DtxR is 9 A. D8M binds only a single Mn2+ per monomer, owing to alteration of the metal-binding site. The sole retained metal site adopts pseudo-hexacoordinate geometry rather than the pseudo-heptacoordinate geometry of the MntR metal sites.
- Department of Chemistry, Reed College, 3203 SE Woodstock Blvd., Portland, Oregon 97202, USA. glasfeld@reed.edu
Organizational Affiliation: