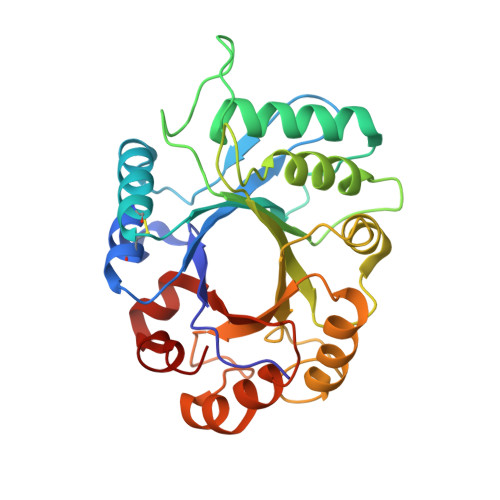
Structural analysis of xylanase inhibitor protein I (XIP-I), a proteinaceous xylanase inhibitor from wheat (Triticum aestivum, var. Soisson).
Payan, F., Flatman, R., Porciero, S., Williamson, G., Juge, N., Roussel, A.(2003) Biochem J 372: 399-405
- PubMed: 12617724
- DOI: https://doi.org/10.1042/BJ20021802
- Primary Citation Related Structures:
1OM0 - PubMed Abstract:
A novel class of proteinaceous inhibitors exhibiting specificity towards microbial xylanases has recently been discovered in cereals. The three-dimensional structure of xylanase inhibitor protein I (XIP-I) from wheat (Triticum aestivum, var. Soisson) was determined by X-ray crystallography at 1.8 A (1 A=0.1 nm) resolution. The inhibitor possesses a (beta/alpha)(8) barrel fold and has structural features typical of glycoside hydrolase family 18, namely two consensus regions, approximately corresponding to the third and fourth barrel strands, and two non-proline cis -peptide bonds, Ser(36)-Phe and Trp(256)-Asp (in XIP-I numbering). However, detailed structural analysis of XIP-I revealed several differences in the region homologous with the active site of chitinases. The catalytic glutamic acid residue of family 18 chitinases [Glu(127) in hevamine, a chitinase/lysozyme from the rubber tree (Hevea brasiliensis)] is conserved in the structure of the inhibitor (Glu(128)), but its side chain is fully engaged in salt bridges with two neighbouring arginine residues. Gly(81), located in subsite -1 of hevamine, where the reaction intermediate is formed, is replaced by Tyr(80) in XIP-I. The tyrosine side chain fills the subsite area and makes a strong hydrogen bond with the side chain of Glu(190) located at the opposite side of the cleft, preventing access of the substrate to the catalytic glutamic acid. The structural differences in the inhibitor cleft structure probably account for the lack of activity of XIP-I towards chitin.
- Architecture et Fonction des Macromolécules Biologiques, UMR6098, CNRS and Universities Aix-Marseille I and II, 31 chemin Joseph Aiguier, France. fran@afmb.cnrs-mrs.fr
Organizational Affiliation: