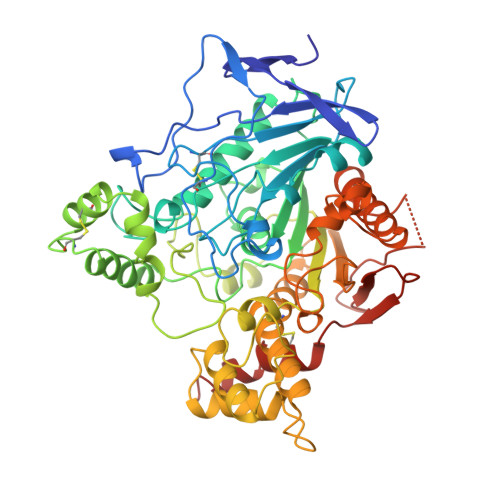
"Back door" opening implied by the crystal structure of a carbamoylated acetylcholinesterase.
Bartolucci, C., Perola, E., Cellai, L., Brufani, M., Lamba, D.(1999) Biochemistry 38: 5714-5719
- PubMed: 10231521 Search on PubMed
- DOI: https://doi.org/10.1021/bi982723p
- Primary Citation Related Structures:
1OCE - PubMed Abstract:
The crystal structure of Torpedo californica (Tc) acetylcholinesterase (AChE) carbamoylated by the physostigmine analogue 8-(cis-2,6-dimethylmorpholino)octylcarbamoyleseroline (MF268) is reported at 2.7 A resolution. In the X-ray structure, the dimethylmorpholinooctylcarbamic moiety of MF268 is covalently bound to the catalytic serine, which is located at the bottom of a long and narrow gorge. The alkyl chain of the inhibitor fills the upper part of the gorge, blocking the entrance of the active site. This prevents eseroline, the leaving group of the carbamoylation process, from exiting through this path. Surprisingly, the relatively bulky eseroline is not found in the crystal structure, thus implying the existence of an alternative route for its clearance. This represents indirect evidence that a "back door" opening may occur and shows that the release of products via a "back door" is a likely alternative for this enzyme. However, its relevance as far as the mechanism of substrate hydrolysis is concerned needs to be established. This study suggests that the use of properly designed acylating inhibitors, which can block the entrance of catalytic sites, may be exploited as a general approach for investigating the existence of "back doors" for the clearance of products.
- Istituto di Strutturistica Chimica "G. Giacomello", CNR, Rome, Italy.
Organizational Affiliation: