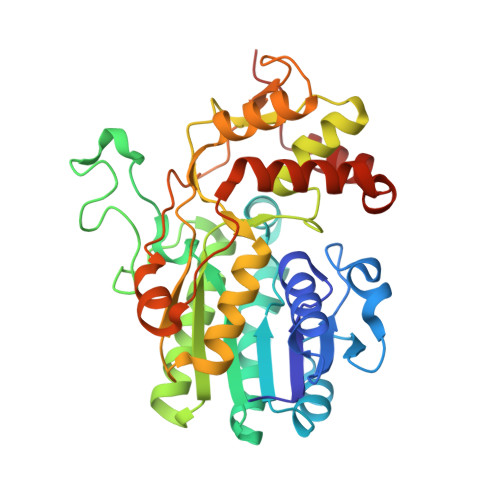
The Structure of Nadh in the Enzyme Dtdp-D-Glucose Dehydratase (Rmlb)
Beis, K., Allard, S.T., Hegeman, A.D., Murshudov, G., Philp, D., Naismith, J.H.(2003) J Am Chem Soc 125: 11872
- PubMed: 14505409 Search on PubMed
- DOI: https://doi.org/10.1021/ja035796r
- Primary Citation Related Structures:
1OC2 - PubMed Abstract:
The structure of Streptococcus suis serotype type 2 dTDP-d-glucose 4,6-dehydratase (RmlB) has been determined to 1.5 A resolution with its nicotinamide coenzyme and substrate analogue dTDP-xylose bound in an abortive complex. During enzyme turnover, NAD(+) abstracts a hydride from the C4' atom of dTDP-glucose-forming NADH. After elimination of water, hydride is then transferred back to the C6' atom of dTDP-4-keto-5,6-glucosene-regenerating NAD(+). Single-crystal spectroscopic studies unambiguously show that the coenzyme has been trapped as NADH in the crystal. Electron density clearly demonstrates that in contrast to native structures of RmlB where a flat nicotinamide ring is observed, the dihydropyridine ring of the reduced cofactor in this complex is found as a boat. The si face, from which the pro-S hydride is transferred, has a concave surface. Ab initio electronic structure calculations demonstrate that the presence of an internal hydrogen bond, between the amide NH on the nicotinamide ring and one of the oxygen atoms on a phosphate group, stabilizes this distorted conformation. Additionally, calculations show that the hydride donor ability of NADH is influenced by the degree of bending in the ring and may be influenced by an active-site tyrosine residue (Tyr 161). These results demonstrate the ability of dehydratase enzymes to fine-tune the redox potential of NADH through conformational changes in the nicotinamide ring.
- Centre for Biomolecular Sciences, University of St. Andrews, North Haugh, St. Andrews, Fife KY16 9ST, United Kingdom.
Organizational Affiliation: