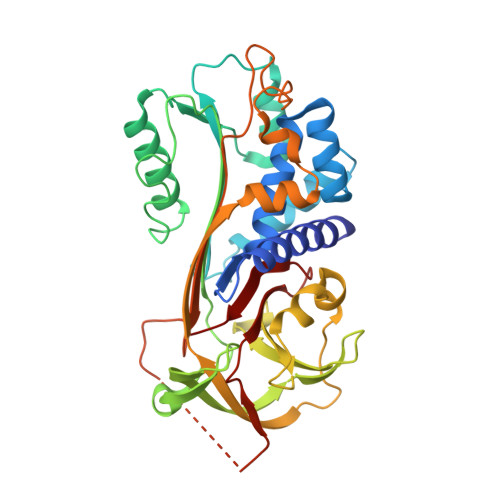
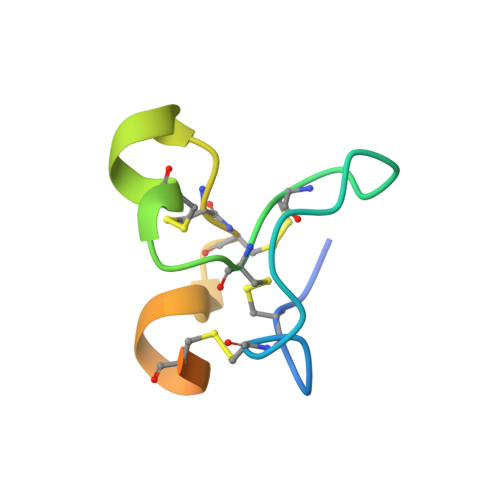
How Vitronectin Binds Pai-1 to Modulate Fibrinolysis and Cell Migration
Zhou, A., Huntington, J.A., Pannu, N.S., Carrell, R.W., Read, R.J.(2003) Nat Struct Biol 10: 541
- PubMed: 12808446 Search on PubMed
- DOI: https://doi.org/10.1038/nsb943
- Primary Citation Related Structures:
1OC0 - PubMed Abstract:
The interaction of the plasma protein vitronectin with plasminogen activator inhibitor-1 (PAI-1) is central to human health. Vitronectin binding extends the lifetime of active PAI-1, which controls hemostasis by inhibiting fibrinolysis and has also been implicated in angiogenesis. The PAI-1-vitronectin binding interaction also affects cell adhesion and motility. For these reasons, elevated PAI-1 activities are associated both with coronary thrombosis and with a poor prognosis in many cancers. Here we show the crystal structure at a resolution of 2.3 A of the complex of the somatomedin B domain of vitronectin with PAI-1. The structure of the complex explains how vitronectin binds to and stabilizes the active conformation of PAI-1. It also explains the tissue effects of PAI-1, as PAI-1 competes for and sterically blocks the interaction of vitronectin with cell surface receptors and integrins. Structural understanding of the essential biological roles of the interaction between PAI-1 and vitronectin opens the prospect of specifically designed blocking agents for the prevention of thrombosis and treatment of cancer.
- Department of Haematology, University of Cambridge, Cambridge Institute for Medical Research, Wellcome Trust/MRC Building, Hills Road, Cambridge CB2 2XY, UK.
Organizational Affiliation: