Crystallization and preliminary crystallographic study of the peptidoglycan-associated lipoprotein from Escherichia coli.
Abergel, C., Walburger, A., Chenivesse, S., Lazdunski, C.(2001) Acta Crystallogr D Biol Crystallogr 57: 317-319
- PubMed: 11173492
- DOI: https://doi.org/10.1107/s0907444900019739
- Primary Citation Related Structures:
1OAP - PubMed Abstract:
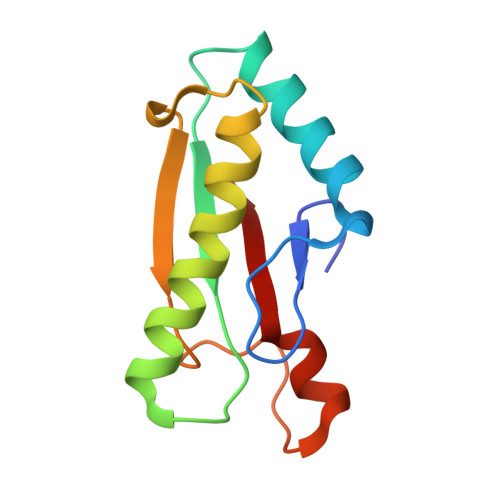
The peptidoglycan-associated lipoprotein (Pal) from Escherichia coli is part of the Tol--Pal multiprotein complex used by group A colicins to penetrate and kill cells. Pal homologues are found in many Gram-negative bacteria and the Tol--Pal system is thought to play a role in bacterial envelope integrity. The Pal protein comprises 152 amino acids. Crystals of the C-terminal 109-amino-acid fragment of the Pal protein have been produced. The crystals belong to the tetragonal space group I4(1), with unit-cell parameters a = b = 89.3, c = 67.2 A. There are two molecules in the asymmetric unit. Frozen crystals diffract to at least 2.8 A resolution using synchrotron radiation. Selenomethionine-substituted truncated Pal protein is currently being produced in order to use multiwavelength anomalous dispersion (MAD) for phasing.
- Information Génétique et Structurale, UMR1889 CNRS-AVENTIS, 31 Chemin Joseph Aiguier, 13402 Marseille CEDEX 20, France. chantal@igs.cnrs-mrs.fr
Organizational Affiliation: