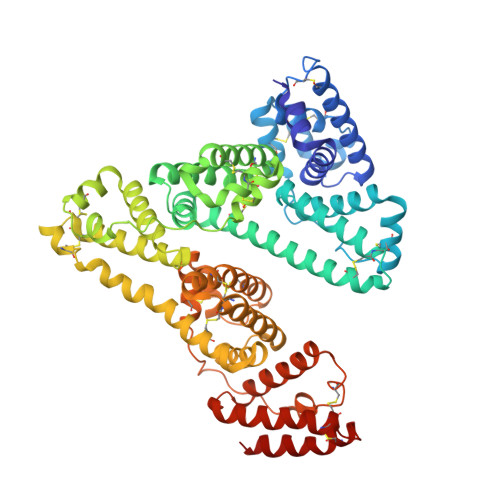
Crystal Structure Analysis of Human Serum Albumin Complexed with Hemin and Fatty Acid
Zunszain, P.A., Ghuman, J., Komatsu, T., Tsuchida, E., Curry, S.(2003) BMC Struct Biol 3: 6
- PubMed: 12846933 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-3-6
- Primary Citation Related Structures:
1O9X - PubMed Abstract:
Human serum albumin (HSA) is an abundant plasma protein that binds a wide variety of hydrophobic ligands including fatty acids, bilirubin, thyroxine and hemin. Although HSA-heme complexes do not bind oxygen reversibly, it may be possible to develop modified HSA proteins or heme groups that will confer this ability on the complex. We present here the crystal structure of a ternary HSA-hemin-myristate complex, formed at a 1:1:4 molar ratio, that contains a single hemin group bound to subdomain IB and myristate bound at six sites. The complex displays a conformation that is intermediate between defatted HSA and HSA-fatty acid complexes; this is likely to be due to low myristate occupancy in the fatty acid binding sites that drive the conformational change. The hemin group is bound within a narrow D-shaped hydrophobic cavity which usually accommodates fatty acid; the hemin propionate groups are coordinated by a triad of basic residues at the pocket entrance. The iron atom in the centre of the hemin is coordinated by Tyr161. The structure of the HSA-hemin-myristate complex (PDB ID 1o9x) reveals the key polar and hydrophobic interactions that determine the hemin-binding specificity of HSA. The details of the hemin-binding environment of HSA provide a structural foundation for efforts to modify the protein and/or the heme molecule in order to engineer complexes that have favourable oxygen-binding properties.
- Department of Biological Sciences, Imperial College London, Room 746 Huxley Building, South Kensington Campus, London SW7 2AZ, United Kingdom. p.zunszain@imperial.ac.uk
Organizational Affiliation: