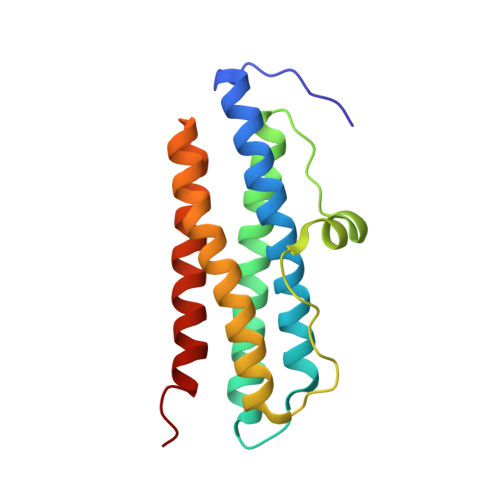
The Dps Protein of Agrobacterium Tumefaciens Does not Bind to DNA But Protects It Toward Oxidative Cleavage: X-Ray Crystal Structure, Iron Binding, and Hydroxyl-Radical Scavenging Properties
Ceci, P., Ilari, A., Falvo, E., Chiancone, E.(2003) J Biological Chem 278: 20319
- PubMed: 12660233
- DOI: https://doi.org/10.1074/jbc.M302114200
- Primary Citation Related Structures:
1O9R - PubMed Abstract:
Agrobacterium tumefaciens Dps (DNA-binding proteins from starved cells), encoded by the dps gene located on the circular chromosome of this plant pathogen, was cloned, and its structural and functional properties were determined in vitro. In Escherichia coli Dps, the family prototype, the DNA binding properties are thought to be associated with the presence of the lysine-containing N-terminal tail that extends from the protein surface into the solvent. The x-ray crystal structure of A. tumefaciens Dps shows that the positively charged N-terminal tail, which is 11 amino acids shorter than in the E. coli protein, is blocked onto the protein surface. This feature accounts for the lack of interaction with DNA. The intersubunit ferroxidase center characteristic of Dps proteins is conserved and confers to the A. tumefaciens protein a ferritin-like activity that manifests itself in the capacity to oxidize and incorporate iron in the internal cavity and to release it after reduction. In turn, sequestration of Fe(II) correlates with the capacity of A. tumefaciens Dps to reduce the production of hydroxyl radicals from H2O2 through Fenton chemistry. These data demonstrate conclusively that DNA protection from oxidative damage in vitro does not require formation of a Dps-DNA complex. In vivo, the hydroxyl radical scavenging activity of A. tumefaciens Dps may be envisaged to act in concert with catalase A to counteract the toxic effect of H2O2, the major component of the plant defense system when challenged by the bacterium.
- Consiglio Nazionale delle Ricerche Institute of Molecular Biology and Pathology, Department of Biochemical Sciences A. Rossi-Fanelli, University of Rome La Sapienza, Italy.
Organizational Affiliation: