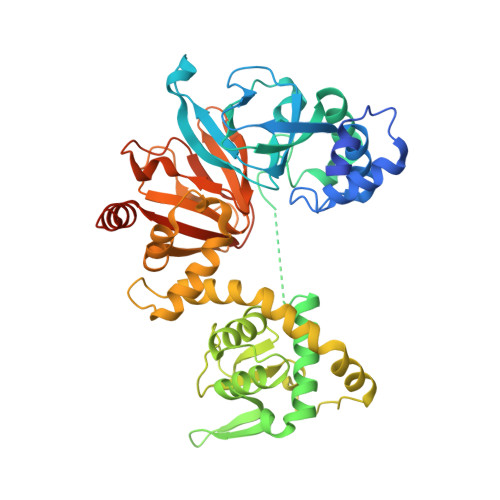
Structure and Regulation of the Camp-Binding Domains of Epac2
Rehmann, H., Prakash, B., Wolf, E., Rueppel, A., De Rooij, J., Bos, J.L., Wittinghofer, A.(2002) Nat Struct Biol 10: 26
- PubMed: 12469113
- DOI: https://doi.org/10.1038/nsb878
- Primary Citation of Related Structures:
1O7F - PubMed Abstract:
Cyclic adenosine monophosphate (cAMP) is a universal second messenger that, in eukaryotes, was believed to act only on cAMP-dependent protein kinase A (PKA) and cyclic nucleotide-regulated ion channels. Recently, guanine nucleotide exchange factors specific for the small GTP-binding proteins Rap1 and Rap2 (Epacs) were described, which are also activated directly by cAMP. Here, we have determined the three-dimensional structure of the regulatory domain of Epac2, which consists of two cyclic nucleotide monophosphate (cNMP)-binding domains and one DEP (Dishevelled, Egl, Pleckstrin) domain. This is the first structure of a cNMP-binding domain in the absence of ligand, and comparison with previous structures, sequence alignment and biochemical experiments allow us to delineate a mechanism for cyclic nucleotide-mediated conformational change and activation that is most likely conserved for all cNMP-regulated proteins. We identify a hinge region that couples cAMP binding to a conformational change of the C-terminal regions. Mutations in the hinge of Epac can uncouple cAMP binding from its exchange activity.
- Max-Planck Institut für Molekulare Physiologie, Otto Hahn Strasse 11, D-44227, Dortmund, Germany.
Organizational Affiliation: