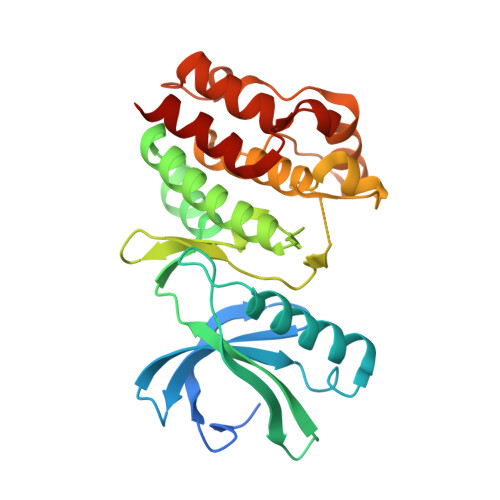
Crystal Structure of the Catalytic Domain of the Pknb Serine/Threonine Kinase from Mycobacterium Tuberculosis
Ortiz-Lombardia, M., Pompeo, F., Boitel, B., Alzari, P.M.(2003) J Biological Chem 278: 13094
- PubMed: 12551895 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M300660200
- Primary Citation Related Structures:
1O6Y - PubMed Abstract:
With the advent of the sequencing programs of prokaryotic genomes, many examples of the presence of serine/threonine protein kinases in these organisms have been identified. Moreover, these kinases could be classified as homologues of those belonging to the well characterized superfamily of the eukaryotic serine/threonine and tyrosine kinases. Eleven such kinases were recognized in the genome of Mycobacterium tuberculosis. Here we report the crystal structure of an active form of PknB, one of the four M. tuberculosis kinases that are conserved in the downsized genome of Mycobacterium leprae and are therefore presumed to play an important role in the processes that regulate the complex life cycle of mycobacteria. Our structure confirms again the extraordinary conservation of the protein kinase fold and constitutes a landmark that extends this conservation across the evolutionary distance between high eukaryotes and eubacteria. The structure of PknB, in complex with a nucleotide triphosphate analog, reveals an enzyme in the active state with an unprecedented arrangement of the Gly-rich loop associated with a new conformation of the nucleotide gamma-phosphoryl group. It presents as well a partially disordered activation loop, suggesting an induced fit mode of binding for the so far unknown substrates of this kinase or for some modulating factor(s).
- Unité de Biochimie Structurale, URA 2185 CNRS, Institut Pasteur, 25, rue du Dr. Roux, 75724 Paris, cedex 15, France.
Organizational Affiliation: