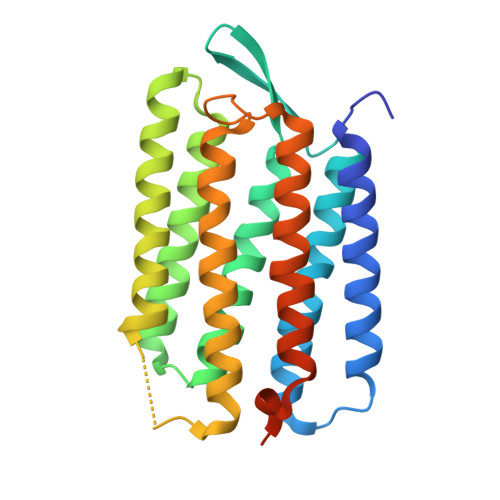
Mechanism of proton transport in bacteriorhodopsin from crystallographic structures of the K, L, M1, M2, and M2' intermediates of the photocycle.
Lanyi, J.K., Schobert, B.(2003) J Mol Biology 328: 439-450
- PubMed: 12691752 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00263-8
- Primary Citation Related Structures:
1O0A - PubMed Abstract:
We produced the L intermediate of the photocycle in a bacteriorhodopsin crystal in photo-stationary state at 170 K with red laser illumination at 60% occupancy, and determined its structure to 1.62 A resolution. With this model, high-resolution structural information is available for the initial bacteriorhodopsin, as well as the first five states in the transport cycle. These states involve photo-isomerization of the retinal and its initial configurational changes, deprotonation of the retinal Schiff base and the coupled release of a proton to the extracellular membrane surface, and the switch event that allows reprotonation of the Schiff base from the cytoplasmic side. The six structural models describe the transformations of the retinal and its interaction with water 402, Asp85, and Asp212 in atomic detail, as well as the displacements of functional residues farther from the Schiff base. The changes provide rationales for how relaxation of the distorted retinal causes movements of water and protein atoms that result in vectorial proton transfers to and from the Schiff base.
- Department of Physiology and Biophysics, University of California, 346D Medical Science I, Irvine, CA 92697, USA. jlanyi@orion.oac.uci.edu
Organizational Affiliation: