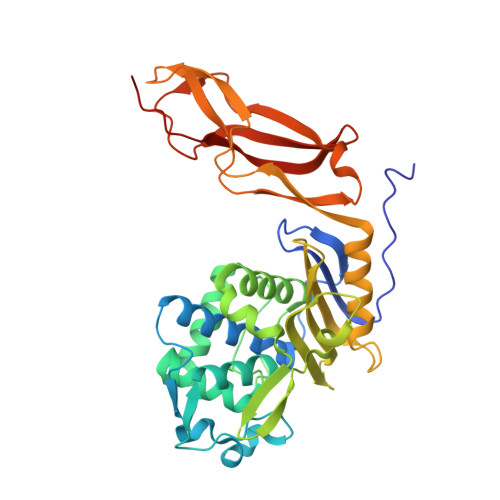
Crystal structure of wild-type penicillin-binding protein 5 from Escherichia coli: implications for deacylation of the acyl-enzyme complex.
Nicholas, R.A., Krings, S., Tomberg, J., Nicola, G., Davies, C.(2003) J Biological Chem 278: 52826-52833
- PubMed: 14555648 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M310177200
- Primary Citation Related Structures:
1NJ4, 1NZO - PubMed Abstract:
Penicillin-binding protein 5 (PBP 5) of Escherichia coli functions as a d-alanine carboxypeptidase (CPase), cleaving d-alanine from the C terminus of cell wall peptides. Like all PBPs, PBP 5 forms a covalent acyl-enzyme complex with beta-lactam antibiotics; however, PBP 5 is distinguished by its high rate of deacylation of the acylenzyme complex (t(1/2) approximately 10 min). A Gly105 --> Asp mutation in PBP 5 markedly impairs deacylation with only minor effects on acylation, and abolishes CPase activity. We have determined the three-dimensional structure of a soluble form of wild-type PBP 5 at 1.85-A resolution and have also refined the structure of the G105D mutant form of PBP 5 to 1.9-A resolution. Comparison of the two structures reveals that the major effect of the mutation is to disorder a loop comprising residues 74-90 that sits atop the SXN motif of the active site. Deletion of the 74-90 loop in wild-type PBP 5 markedly diminished the deacylation rate of penicillin G with a minimal impact on acylation, and abolished CPase activity. These effects were very similar to those observed in the G105D mutant, reinforcing the idea that this mutation causes disordering of the 74-90 loop. Mutation of two consecutive serines within this loop, which hydrogen bond to Ser110 and Asn112 in the SXN motif, had marked effects on CPase activity, but not beta-lactam antibiotic binding or hydrolysis. These data suggest a direct role for the SXN motif in deacylation of the acyl-enzyme complex and imply that the functioning of this motif is modulated by the 74-90 loop.
- Department of Pharmacology, University of North Carolina, Chapel Hill, North Carolina 27599-7365, USA. nicholas@med.unc.edu
Organizational Affiliation: