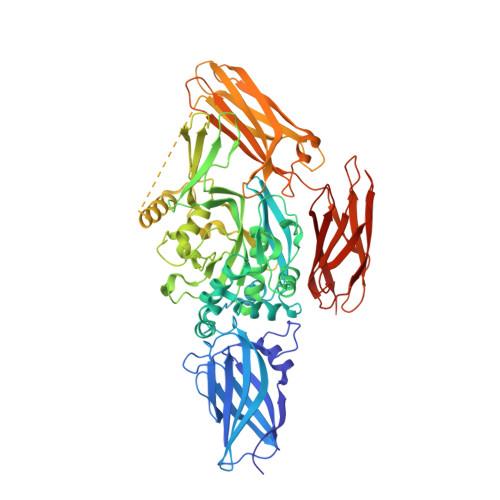
Roles of Calcium Ions in the Activation and Activity of the Transglutaminase 3 Enzyme
Ahvazi, B., Boeshans, K.M., Idler, W., Baxa, U., Steinert, P.M.(2003) J Biological Chem 278: 23834-23841
- PubMed: 12679341
- DOI: https://doi.org/10.1074/jbc.M301162200
- Primary Citation Related Structures:
1NUD, 1NUF, 1NUG - PubMed Abstract:
The transglutaminase 3 enzyme is widely expressed in many tissues including epithelia. We have shown previously that it can bind three Ca2+ ions, which in site one is constitutively bound, while those in sites two and three are acquired during activation and are required for activity. In particular, binding at site three opens a channel through the enzyme and exposes two tryptophan residues near the active site that are thought to be important for enzyme reaction. In this study, we have solved the structures of three more forms of this enzyme by x-ray crystallography in the presence of Ca2+ and/or Mg2+, which provide new insights on the precise contribution of each Ca2+ ion to activation and activity. First, we found that Ca2+ ion in site one can be exchanged with difficulty, and it has a binding affinity of Kd = 0.3 microm (DeltaH = -6.70 +/- 0.52 kcal/mol), which suggests it is important for the stabilization of the enzyme. Site two can be occupied by some lanthanides but only Ca2+ of the Group 2 family of alkali earth metals, and its occupancy are required for activity. Site three can be occupied by some lanthanides, Ca2+,or Mg2+; however, when Mg2+ is present, the enzyme is inactive, and the channel is closed. Thus Ca2+ binding in both sites two and three cooperate in opening the channel. We speculate that manipulation of the channel opening could be controlled by intracellular cation levels. Together, these data have important implications for reaction mechanism of the enzyme: the opening of a channel perhaps controls access to and manipulation of substrates at the active site.
- Laboratory of Skin Biology, the Laboratory of X-ray Crystallography/Office of Science and Technology, National Institute of Health, Bethesda, Maryland 20892-8023, USA. Bijan@discus.niams.nih.gov
Organizational Affiliation: