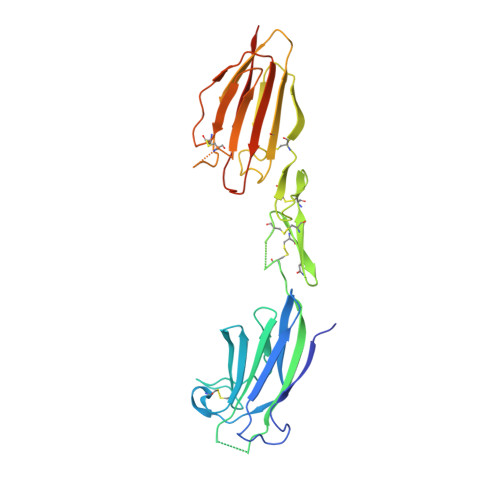
Crystal structure of the CUB1-EGF-CUB2 region of mannose-binding protein associated serine protease-2
Feinberg, H., Uitdehaag, J.C.M., Davies, J.M., Wallis, R., Drickamer, K., Weis, W.I.(2003) EMBO J 22: 2348-2359
- PubMed: 12743029 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg236
- Primary Citation Related Structures:
1NT0 - PubMed Abstract:
Serum mannose-binding proteins (MBPs) are C-type lectins that recognize cell surface carbohydrate structures on pathogens, and trigger killing of these targets by activating the complement pathway. MBPs circulate as a complex with MBP-associated serine proteases (MASPs), which become activated upon engagement of a target cell surface. The minimal functional unit for complement activation is a MASP homodimer bound to two MBP trimeric subunits. MASPs have a modular structure consisting of an N-terminal CUB domain, a Ca(2+)-binding EGF-like domain, a second CUB domain, two complement control protein modules and a C-terminal serine protease domain. The CUB1-EGF-CUB2 region mediates homodimerization and binding to MBP. The crystal structure of the MASP-2 CUB1-EGF-CUB2 dimer reveals an elongated structure with a prominent concave surface that is proposed to be the MBP-binding site. A model of the full six-domain structure and its interaction with MBPs suggests mechanisms by which binding to a target cell transmits conformational changes from MBP to MASP that allow activation of its protease activity.
- Departments of Structural Biology and of Molecular and Cellular Physiology, Stanford University School of Medicine, 299 Campus Drive West, CA 94305-5126, USA.
Organizational Affiliation: