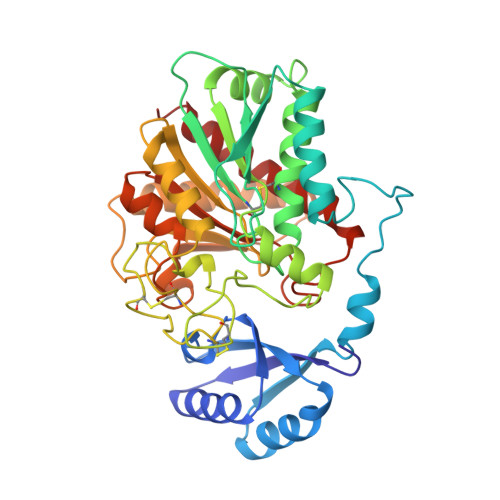
Three-dimensional structure of porcine procarboxypeptidase B: a structural basis of its inactivity.
Coll, M., Guasch, A., Aviles, F.X., Huber, R.(1991) EMBO J 10: 1-9
- PubMed: 1989878 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1991.tb07914.x
- Primary Citation Related Structures:
1NSA - PubMed Abstract:
Procarboxypeptidase B is converted to enzymatically active carboxypeptidase B by limited proteolysis catalysed by trypsin, removing the long N-terminal activation segment of 95 amino acids. The three-dimensional crystal structure of procarboxypeptidase B from porcine pancreas has been determined at 2.3 A resolution and refined to a crystallographic R-factor of 0.169. The functional determinants of its enzymatic inactivity and of its activation by limited proteolysis have thus been unveiled. The activation segment folds in a globular region with an open sandwich antiparallel-alpha antiparallel-beta topology and in a C terminal alpha-helix which connects it to the enzyme moiety. The globular region (A7-A82) shields the preformed active site, and establishes specific interactions with residues important for substrate recognition. AspA41 forms a salt bridge with Arg145, which in active carboxypeptidase binds the C-terminal carboxyl group of substrate molecules. The connecting region occupies the putative extended substrate binding site. The scissile peptide bond cleaved by trypsin during activation is very exposed. Its cleavage leads to the release of the activation segment and to exposure of the substrate binding site. An open-sandwich folding has been observed in a number of other proteins and protein domains. One of them is the C-terminal fragment of L7/L12, a ribosomal protein from Escherichia coli that displays a topology similar to the activation domain of procarboxypeptidase.
- Max-Planck-Institut für Biochemie, Martinsried bei München, FRG.
Organizational Affiliation: