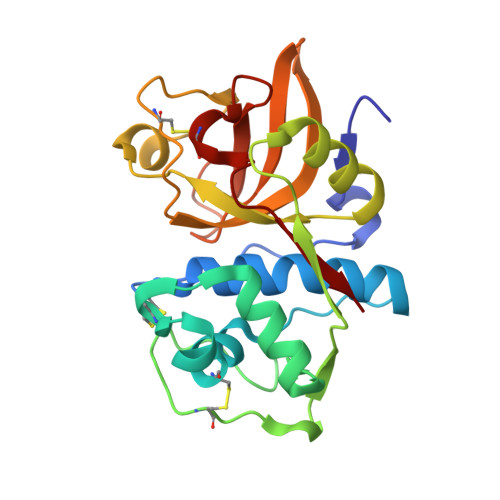
Specificity determinants of human cathepsin s revealed by crystal structures of complexes.
Pauly, T.A., Sulea, T., Ammirati, M., Sivaraman, J., Danley, D.E., Griffor, M.C., Kamath, A.V., Wang, I.K., Laird, E.R., Seddon, A.P., Menard, R., Cygler, M., Rath, V.L.(2003) Biochemistry 42: 3203-3213
- PubMed: 12641451 Search on PubMed
- DOI: https://doi.org/10.1021/bi027308i
- Primary Citation Related Structures:
1NPZ, 1NQC - PubMed Abstract:
Cathepsin S, a lysosomal cysteine protease of the papain superfamily, has been implicated in the preparation of MHC class II alphabeta-heterodimers for antigen presentation to CD4+ T lymphocytes and is considered a potential target for autoimmune-disease therapy. Selective inhibition of this enzyme may be therapeutically useful for attenuating the hyperimmune responses in a number of disorders. We determined the three-dimensional crystal structures of human cathepsin S in complex with potent covalent inhibitors, the aldehyde inhibitor 4-morpholinecarbonyl-Phe-(S-benzyl)Cys-Psi(CH=O), and the vinyl sulfone irreversible inhibitor 4-morpholinecarbonyl-Leu-Hph-Psi(CH=CH-SO(2)-phenyl) at resolutions of 1.8 and 2.0 A, respectively. In the structure of the cathepsin S-aldehyde complex, the tetrahedral thiohemiacetal adduct favors the S-configuration, in which the oxygen atom interacts with the imidazole group of the active site His164 rather than with the oxyanion hole. The present structures provide a detailed map of noncovalent intermolecular interactions established in the substrate-binding subsites S3 to S1' of cathepsin S. In the S2 pocket, which is the binding affinity hot spot of cathepsin S, the Phe211 side chain can assume two stable conformations that accommodate either the P2-Leu or a bulkier P2-Phe side chain. This structural plasticity of the S2 pocket in cathepsin S explains the selective inhibition of cathepsin S over cathepsin K afforded by inhibitors with the P2-Phe side chain. Comparison with the structures of cathepsins K, V, and L allows delineation of local intermolecular contacts that are unique to cathepsin S.
- Exploratory Medicinal Sciences and Computational Chemistry, Groton Laboratories, Pfizer Global Research and Development, Eastern Point Road, Groton, Connecticut 06340, USA.
Organizational Affiliation: