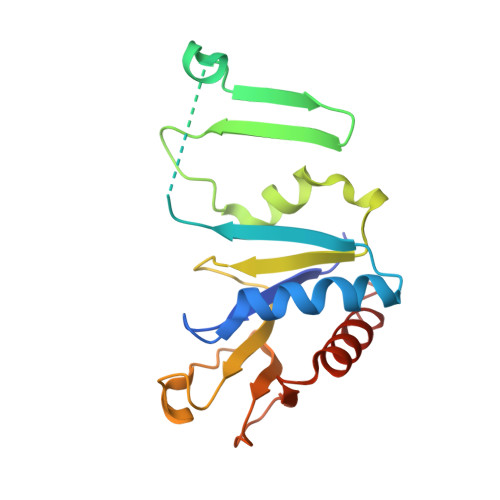
Insight into the role of Escherichia coli MobB in Molybdenum cofactor biosynthesis based on the high resolution crystal structure
McLuskey, K., Harrison, J.A., Schuttelkopf, A.W., Boxer, D.H., Hunter, W.N.(2003) J Biological Chem 278: 23706-23713
- PubMed: 12682065 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M301485200
- Primary Citation Related Structures:
1NP6 - PubMed Abstract:
Two proteins, which are co-transcribed in Escherichia coli (MobA and MobB), are involved in the attachment of a nucleotide moiety to the molybdenum cofactor to form active molybdopterin guanine dinucleotide. Although not essential for this process, the dimeric MobB increases the activation of molybdoenzymes, incorporating this cofactor by a mechanism that is not understood. The structure of MobB has been elucidated in two crystal forms, one of which has provided a model at 1.9-A resolution with Rwork and Rfree values of 21.5 and 28.7%, respectively. The MobB subunit displays an alpha/beta-fold arranged into a major and minor domain, the latter of which is inserted between the major and minor domains of the partner subunit, creating an elongated dimer constructed around a 16-stranded beta-sheet. Structural homologues have been identified, and they include a number of nucleotide-binding proteins. Comparisons indicate that although the phosphate-binding regions are highly conserved, MobB lacks the elements of structure required to interact with and efficiently bind a nucleotide base. In the present structure, a sulfate is bound to the Walker A phosphate-binding motif of MobB. The possibility that MobB forms a complex with the nucleotide-binding MobA, the protein with which it is co-transcribed, is explored, and modeling suggests that such a MobA:MobB complex is feasible. This hypothesis is supported by recent biochemical evidence indicating that MobB interacts with several proteins involved in various stages of molybdenum cofactor biosynthesis including MobA. We propose therefore that MobB is an adapter protein that acts in concert with MobA to achieve the efficient biosynthesis and utilization of molybdopterin guanine dinucleotide.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: