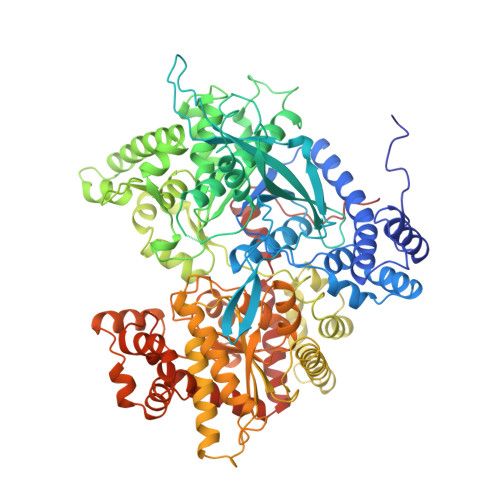
Ternary complex crystal structures of glycogen phosphorylase with the transition state analogue nojirimycin tetrazole and phosphate in the T and R states.
Mitchell, E.P., Withers, S.G., Ermert, P., Vasella, A.T., Garman, E.F., Oikonomakos, N.G., Johnson, L.N.(1996) Biochemistry 35: 7341-7355
- PubMed: 8652510
- DOI: https://doi.org/10.1021/bi960072w
- Primary Citation Related Structures:
1NOI, 1NOJ, 1NOK - PubMed Abstract:
Catalysis by glycogen phosphorylase involves a mechanism in which binding of one substrate tightens the binding of the other substrate to produce a productive ternary enzyme-substrate complex. In this work the molecular basis for this synergism is probed in crystallographic studies on ternary complexes in which the glucosyl component is substituted by the putative transition state analogue nojirimycin tetrazole, a compound which has been established previously as a transition state analogue inhibitor for a number of glycosidases. Kinetic studies with glycogen phosphorylase showed that nojirimycin tetrazole is a competitive inhibitor with respect to glucose 1-phosphate and uncompetitive with respect to phosphate. Ki values for the phosphorylase-AMP-glycogen complex and the phosphorylase-AMP-glycogen-phosphate complexes are 700 microM and 53 microM, respectively, indicating that by itself norjirimycin tetrazole has poor affinity for glycogen phosphorylase but that phosphate substantially improves the binding of norjirimycin tetrazole. X-ray crystallographic binding studies to 2.4 A resolution with T state phosphorylase b crystals showed that nojirimycin tetrazole binds at the catalytic site and promotes the binding of phosphate through direct interactions. Phosphate binding is accompanied by conformational changes that bring a crucial arginine (Arg569) into the catalytic site. The positions of the phosphate oxygens were definitively established in X-ray crystallographic binding experiments at 100 K to 1.7 A resolution using synchrotron radiation. X-ray crystallographic binding studies at 2.5 A resolution with R state glycogen phosphorylase crystals showed that the protein atoms and water molecules in contact with the nojirimycin tetrazole and the phosphate are similar to those in the T state. In both T and R states the phosphate ion is within hydrogen-bonding distance of the cofactor pyridoxal 5'-phosphate group and in ionic contact with the N-1 atom of the tetrazole ring. The results are consistent with previous time-resolved structural studies on complexes with heptenitol and phosphate. The structural and kinetic results suggest that nojirimycin tetrazole in combination with phosphate exhibits properties consistent with a transition state analogue and demonstrate how one promotes the binding of the other.
- Laboratory of Molecular Biophysics, University of Oxford, UK.
Organizational Affiliation: