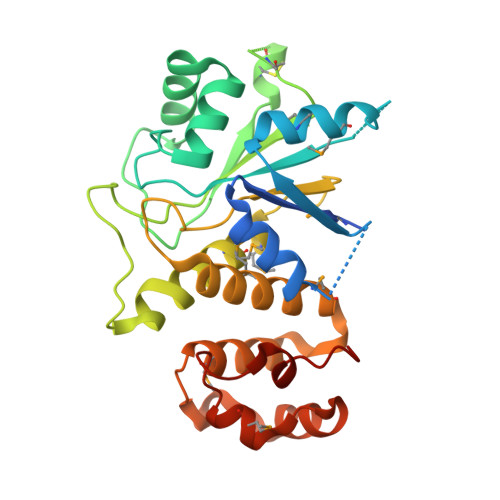
Structure and mechanism of ADP-ribose-1''-monophosphatase (Appr-1''-pase), a ubiquitous cellular processing enzyme
Kumaran, D., Eswaramoorthy, S., Studier, F.W., Swaminathan, S.(2005) Protein Sci 14: 719-726
- PubMed: 15722447 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.041132005
- Primary Citation Related Structures:
1NJR, 1TXZ - PubMed Abstract:
Appr-1''-pase, an important and ubiquitous cellular processing enzyme involved in the tRNA splicing pathway, catalyzes the conversion of ADP-ribose-1''monophosphate (Appr-1''-p) to ADP-ribose. The structures of the native enzyme from the yeast and its complex with ADP-ribose were determined to 1.9 A and 2.05 A, respectively. Analysis of the three-dimensional structure of this protein, selected as a target in a structural genomics project, reveals its putative function and provides clues to the catalytic mechanism. The structure of the 284-amino acid protein shows a two-domain architecture consisting of a three-layer alphabetaalpha sandwich N-terminal domain connected to a small C-terminal helical domain. The structure of Appr-1''-pase in complex with the product, ADP-ribose, reveals an active-site water molecule poised for nucleophilic attack on the terminal phosphate group. Loop-region residues Asn 80, Asp 90, and His 145 may form a catalytic triad.
- Biology Department, Brookhaven National Laboratory, Upton, NY 11973, USA.
Organizational Affiliation: