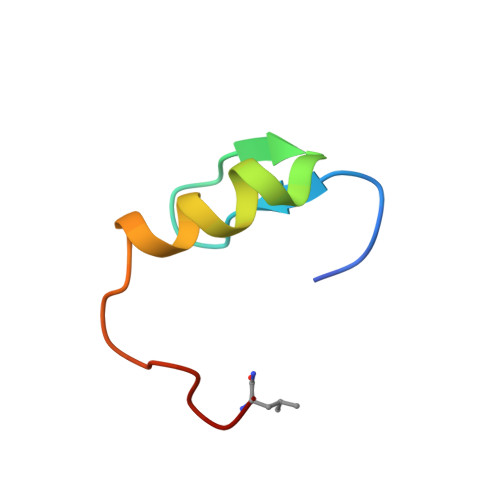
NMR Structure of the Single QALGGH Zinc Finger Domain from the Arabidopsis thaliana SUPERMAN Protein.
Isernia, C., Bucci, E., Leone, M., Zaccaro, L., Di Lello, P., Digilio, G., Esposito, S., Saviano, M., Di Blasio, B., Pedone, C., Pedone, P.V., Fattorusso, R.(2003) Chembiochem 4: 171-180
- PubMed: 12616630 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.200390028
- Primary Citation Related Structures:
1NJQ - PubMed Abstract:
Zinc finger domains of the classical type represent the most abundant DNA binding domains in eukaryotic transcription factors. Plant proteins contain from one to four zinc finger domains, which are characterized by high conservation of the sequence QALGGH, shown to be critical for DNA-binding activity. The Arabidopsis thaliana SUPERMAN protein, which contains a single QALGGH zinc finger, is necessary for proper spatial development of reproductive floral tissues and has been shown to specifically bind to DNA. Here, we report the synthesis and UV and NMR spectroscopic structural characterization of a 37 amino acid SUPERMAN region complexed to a Zn(2+) ion (Zn-SUP37) and present the first high-resolution structure of a classical zinc finger domain from a plant protein. The NMR structure of the SUPERMAN zinc finger domain consists of a very well-defined betabetaalpha motif, typical of all other Cys(2)-His(2) zinc fingers structurally characterized. As a consequence, the highly conserved QALGGH sequence is located at the N terminus of the alpha helix. This region of the domain of animal zinc finger proteins consists of hypervariable residues that are responsible for recognizing the DNA bases. Therefore, we propose a peculiar DNA recognition code for the QALGGH zinc finger domain that includes all or some of the amino acid residues at positions -1, 2, and 3 (numbered relative to the N terminus of the helix) and possibly others at the C-terminal end of the recognition helix. This study further confirms that the zinc finger domain, though very simple, is an extremely versatile DNA binding motif.
- Dipartimento di Scienze Ambientali, Seconda Università di Napoli, 81100 Caserta, Italy.
Organizational Affiliation: