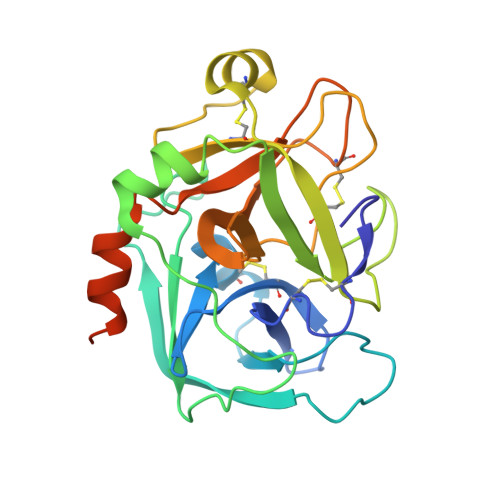
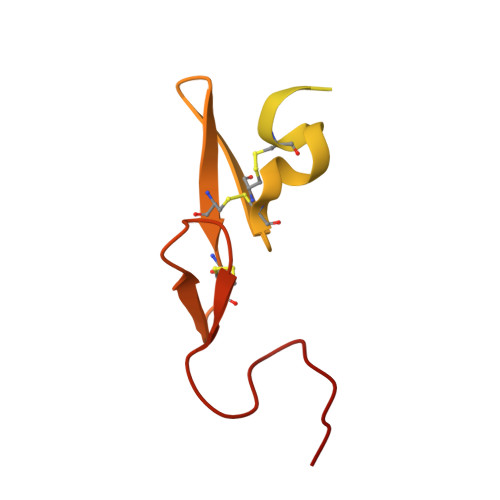
Molecular structures of human Factor Xa complexed with ketopiperazine inhibitors: preference for a neutral group in the S1 pocket.
Maignan, S., Guilloteau, J.P., Choi-Sledeski, Y.M., Becker, M.R., Ewing, W.R., Pauls, H.W., Spada, A.P., Mikol, V.(2003) J Med Chem 46: 685-690
- PubMed: 12593649 Search on PubMed
- DOI: https://doi.org/10.1021/jm0203837
- Primary Citation Related Structures:
1NFU, 1NFW, 1NFX, 1NFY - PubMed Abstract:
The structures of the noncovalent complex of human factor Xa (fXa) with four non-peptide inhibitors containing a central sulfonylpiperazinone scaffold have been determined to about 2.1 A resolution. Highly potent fXa inhibitors containing both neutral groups such as chlorobenzothiophene or chlorothiophene and basic groups such as benzamidine were shown to interact in the S1 pocket through the neutral group whereas the S4 pocket is occupied by the basic moiety. The scaffold comprising the sulfonyl keto piperazine moiety might play a pivotal role in the orientation of substituents, since there is a strong hydrogen bond between Gly219 of fXa and the carbonyl oxygen of the piperazine. This unique "reverse" binding mode is heretofore unreported in fXa and shows that electrostatic interactions in the S1 subsite are not an absolute requirement to maintain high affinity. Selectivity against other serine proteases can be readily explained in light of these structural results. It has opened up new prospects for designing fXa inhibitors with increased oral bioavailability.
- Department of Structural Biology, Aventis Pharma, 13, Quai J. Guesde, F-94403 Vitry/Seine, France.
Organizational Affiliation: