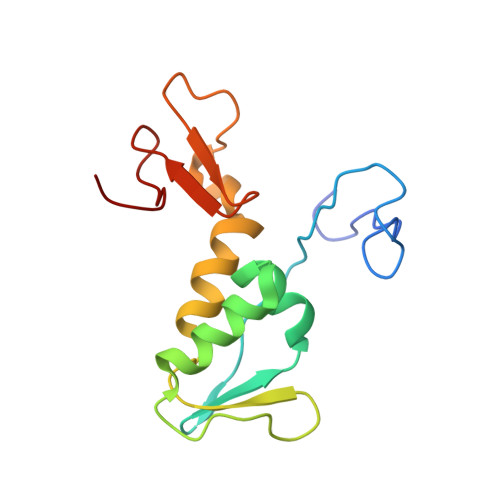
Structure of the archaeal translation initiation factor aIF2beta from Methanobacterium thermoautotrophicum: Implications for translation initiation.
Gutierrez, P., Osborne, M.J., Siddiqui, N., Trempe, J.F., Arrowsmith, C., Gehring, K.(2004) Protein Sci 13: 659-667
- PubMed: 14978306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03506604
- Primary Citation Related Structures:
1NEE - PubMed Abstract:
aIF2 beta is the archaeal homolog of eIF2 beta, a member of the eIF2 heterotrimeric complex, implicated in the delivery of Met-tRNA(i)(Met) to the 40S ribosomal subunit. We have determined the solution structure of the intact beta-subunit of aIF2 from Methanobacterium thermoautotrophicum. aIF2 beta is composed of an unfolded N terminus, a mixed alpha/beta core domain and a C-terminal zinc finger. NMR data shows the two folded domains display restricted mobility with respect to each other. Analysis of the aIF2 gamma structure docked to tRNA allowed the identification of a putative binding site for the beta-subunit in the ternary translation complex. Based on structural similarity and biochemical data, a role for the different secondary structure elements is suggested.
- McGill University, Department of Biochemistry, McIntyre Medical Science Building, 3655 Promenade Sir William Osler, Montréal, Québec H3G 1Y6, Canada.
Organizational Affiliation: