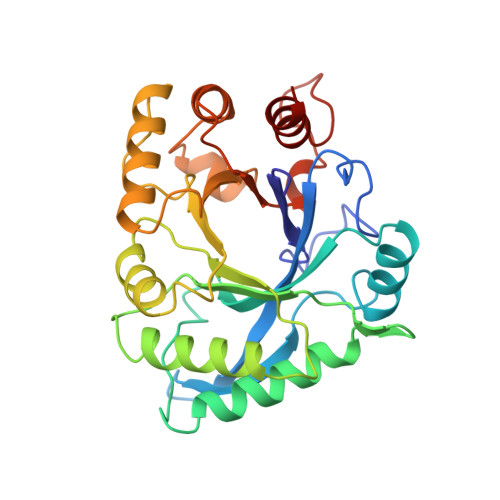
Crystal structure of narbonin at 1.8 A resolution.
Hennig, M., Pfeffer-Hennig, S., Dauter, Z., Wilson, K.S., Schlesier, B., Nong, V.H.(1995) Acta Crystallogr D Biol Crystallogr 51: 177-189
- PubMed: 15299319 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444994009807
- Primary Citation Related Structures:
1NAR - PubMed Abstract:
The three-dimensional structure of narbonin, a seed protein from Vicia narbonensis L, has been determined at 1.8 A resolution. Phase information was obtained by multiple isomorphous replacement and optimized anomalous dispersion. The narbonin structure was initially traced with only 17% amino-acid sequence information and preliminarily refined to a crystallographic R-factor of 16.5%. It is now refined to 15.9% using full sequence information derived from cDNA and after the addition of more solvent molecules. The monomeric molecule of narbonin is an eight-stranded parallel beta-barrel surrounded by alpha-helices in a beta/alpha-topology similar to that first observed in triose phosphate isomerase. Differences exist in the N-terminal part of the polypeptide chain, where the first helix is replaced by a loop and the second beta-strand is followed by an additional antiparallel alpha-sheet placed parallel on top of alpha-helices alpha3 and alpha4. Two short additional secondary structures are present. The first, an alpha-helix, is situated between the seventh beta-strand and the following helix, and the second, which is a 3(10) helix, between the eighth strand and the C-terminal helix. The most striking observation is the lack of a known enzymatic function for narbonin, because all TIM-like structures known so far are enzymes.
- European Molecular Biology Laboratory, Hamburg, Germany.
Organizational Affiliation: