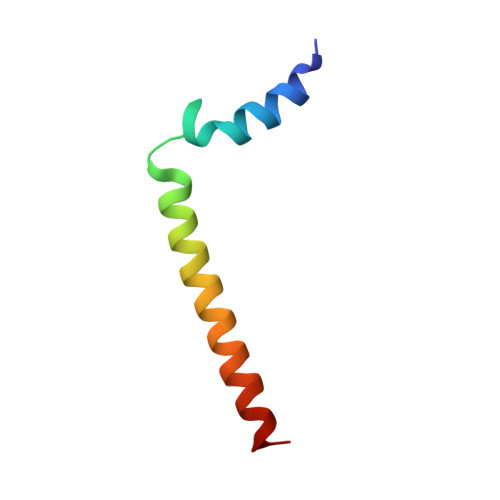
NMR solution structure and topological orientation of monomeric phospholamban in dodecylphosphocholine micelles.
Zamoon, J., Mascioni, A., Thomas, D.D., Veglia, G.(2003) Biophys J 85: 2589-2598
- PubMed: 14507721 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/S0006-3495(03)74681-5
- Primary Citation Related Structures:
1N7L - PubMed Abstract:
Phospholamban is an integral membrane protein that regulates the contractility of cardiac muscle by maintaining cardiomyocyte calcium homeostasis. Abnormalities in association of protein kinase A with PLB have recently been linked to human heart failure, where a single mutation is responsible for dilated cardiomyopathy. To date, a high-resolution structure of phospholamban in a lipid environment has been elusive. Here, we describe the first structure of recombinant, monomeric, biologically active phospholamban in lipid-mimicking dodecylphosphocholine micelles as determined by multidimensional NMR experiments. The overall structure of phospholamban is "L-shaped" with the hydrophobic domain approximately perpendicular to the cytoplasmic portion. This is in agreement with our previously published solid-state NMR data. In addition, there are two striking discrepancies between our structure and those reported previously for synthetic phospholamban in organic solvents: a), in our structure, the orientation of the cytoplasmic helix is consistent with the amphipathic nature of these residues; and b), within the hydrophobic helix, residues are positioned on two discrete faces of the helix as consistent with their functional roles ascribed by mutagenesis. This topology renders the two phosphorylation sites, Ser-16 and Thr-17, more accessible to kinases.
- Department of Biochemistry, Molecular Biology, and Biophysics, and Department of Chemistry, University of Minnesota, Minneapolis, Minnesota 55455, USA.
Organizational Affiliation: