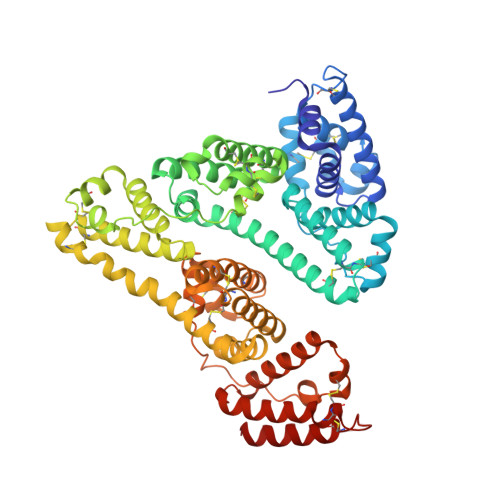
The Atomic Structure of Human Methemalbumin at 1.9 A
Wardell, M., Wang, Z., Ho, J.X., Robert, J., Ruker, F., Ruble, J., Carter, D.C.(2002) Biochem Biophys Res Commun 291: 813-819
- PubMed: 11866438 Search on PubMed
- DOI: https://doi.org/10.1006/bbrc.2002.6540
- Primary Citation Related Structures:
1N5U - PubMed Abstract:
The high resolution structure of hemalbumin was determined by single crystal X-ray diffraction to a resolution of 1.9 A. The structure revealed the protoporphyrin IX bound to a single site within a hydrophobic cavity in subdomain IB, one of the principal binding sites for long chain fatty acid. The iron is penta coordinated with the fifth ligand comprised of the hydroxyl oxygen of Tyr-161 (phenolic oxygen to heme plane distance: 2.73 A) in an otherwise completely hydrophobic pocket. The heme propionic acid residues form salt bridges with His-142 and Lys-190, which together with a series of hydrophobic interactions, enclose and secure the heme within the IB helical motif. A detailed discussion of the structure together with its implications for the development of potential blood substitutes is presented.
- New Century Pharmaceuticals, Inc., 895 Martin Road, Huntsville, Alabama 35824, USA.
Organizational Affiliation: