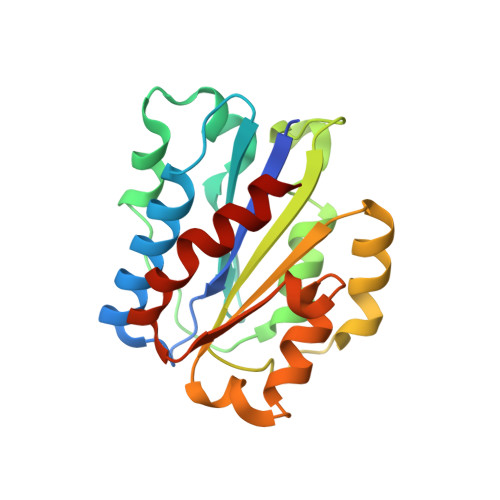
Structure and allosteric regulation of the alpha X beta 2 integrin I domain.
Vorup-Jensen, T., Ostermeier, C., Shimaoka, M., Hommel, U., Springer, T.A.(2003) Proc Natl Acad Sci U S A 100: 1873-1878
- PubMed: 12554829 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0237387100
- Primary Citation Related Structures:
1N3Y - PubMed Abstract:
The integrin alpha X beta 2 (CD11c/CD18, p150,95) binds ligands through the I domain of the alpha X subunit. Ligands include the complement factor fragment iC3b, a key component in the innate immune defense, which, together with the expression of alpha X beta 2 on dendritic cells and on other leukocytes, suggests a role in the immune response. We now report the structure of the alpha X I domain resolved at 1.65 A by x-ray crystallography. To analyze structural requirements for ligand binding we made a mutation in the alpha X I domain C-terminal helix, which increased the affinity for iC3b approximately 200-fold to 2.4 microM compared with the wild-type domain affinity of approximately 400 microM. Gel permeation chromatography supported a conformational change between the wild-type and mutated domains. Conservation of allosteric regulation in the alpha X I domain points to the functional importance of this phenomenon.
- Center for Blood Research, Department of Pathology, Harvard Medical School, 200 Longwood Avenue, Boston, MA 02115, USA.
Organizational Affiliation: