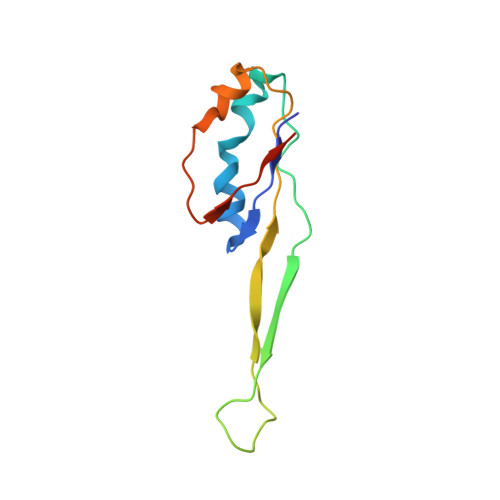
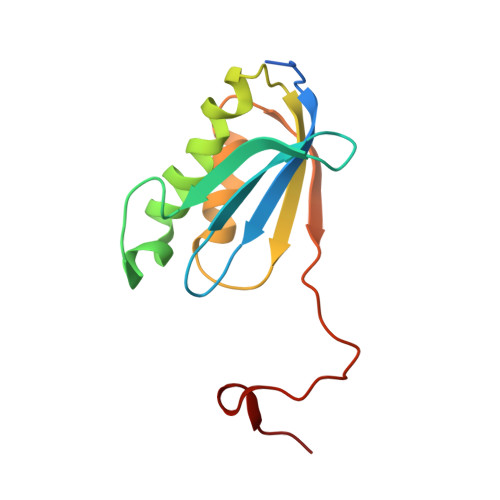
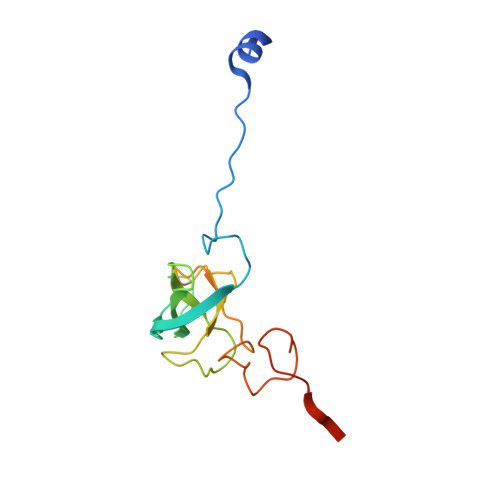
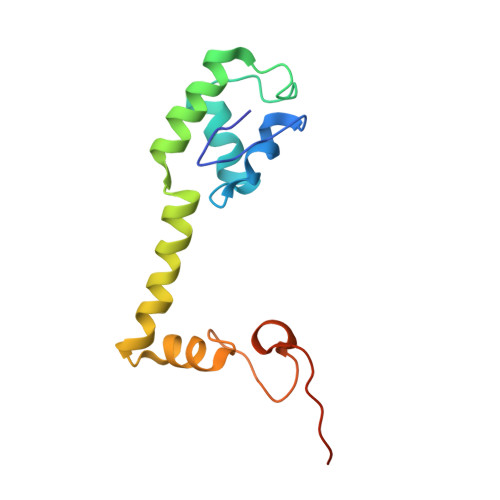
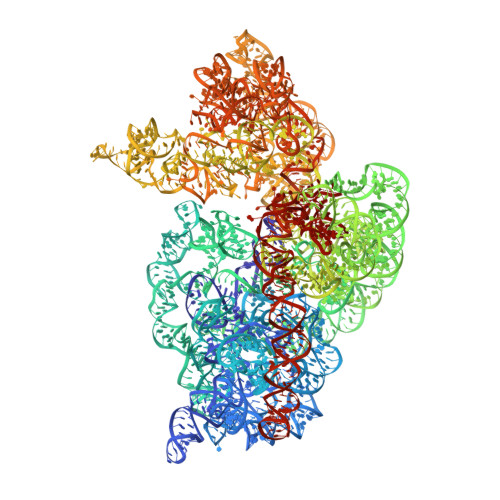
Selection of tRNA by the Ribosome Requires a Transition from an Open to a Closed Form
Ogle, J.M., Murphy IV, F.V., Tarry, M.J., Ramakrishnan, V.(2002) Cell 111: 721-732
- PubMed: 12464183 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(02)01086-3
- Primary Citation Related Structures:
1N32, 1N33, 1N34, 1N36 - PubMed Abstract:
A structural and mechanistic explanation for the selection of tRNAs by the ribosome has been elusive. Here, we report crystal structures of the 30S ribosomal subunit with codon and near-cognate tRNA anticodon stem loops bound at the decoding center and compare affinities of equivalent complexes in solution. In ribosomal interactions with near-cognate tRNA, deviation from Watson-Crick geometry results in uncompensated desolvation of hydrogen-bonding partners at the codon-anticodon minor groove. As a result, the transition to a closed form of the 30S induced by cognate tRNA is unfavorable for near-cognate tRNA unless paromomycin induces part of the rearrangement. We conclude that stabilization of a closed 30S conformation is required for tRNA selection, and thereby structurally rationalize much previous data on translational fidelity.
- MRC Laboratory of Molecular Biology, Hills Road, CB2 2QH, Cambridge, United Kingdom.
Organizational Affiliation: