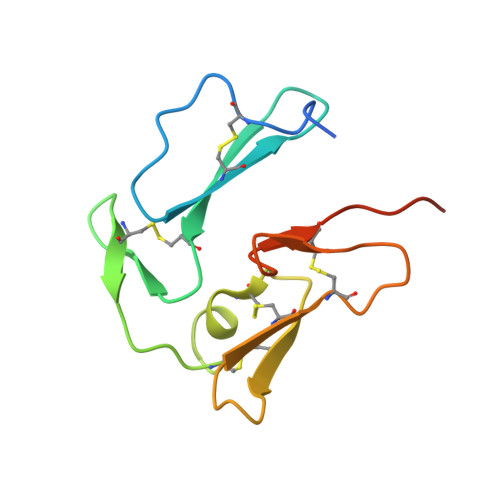
Structure of the C-terminal domains of merozoite surface protein-1 from Plasmodium knowlesi reveals a novel histidine binding site
Garman, S.C., Simcoke, W.N., Stowers, A.W., Garboczi, D.N.(2003) J Biological Chem 278: 7264-7269
- PubMed: 12493733 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M210716200
- Primary Citation Related Structures:
1N1I - PubMed Abstract:
The protozoan parasite Plasmodium causes malaria, with hundreds of millions of cases recorded annually. Protection against malaria infection can be conferred by antibodies against merozoite surface protein (MSP)-1, making it an attractive vaccine candidate. Here we present the structure of the C-terminal domains of MSP-1 (known as MSP-1(19)) from Plasmodium knowlesi. The structure reveals two tightly packed epidermal growth factor-like domains oriented head to tail. In domain 1, the molecule displays a histidine binding site formed primarily by a highly conserved tryptophan. The protein carries a pronounced overall negative charge primarily due to the large number of acidic groups in domain 2. To map protein binding surfaces on MSP-1(19), we have analyzed the crystal contacts in five different crystal environments, revealing that domain 1 is highly preferred in protein-protein interactions. A comparison of MSP-1(19) structures from P. knowlesi, P. cynomolgi, and P. falciparum shows that, although the overall protein folds are similar, the molecules show significant differences in charge distribution. We propose the histidine binding site in domain 1 as a target for inhibitors of protein binding to MSP-1, which might prevent invasion of the merozoite into red blood cells.
- Structural Biology Section, Laboratory of Immunogenetics and Malaria Vaccine Development Unit, Laboratory of Parasitic Diseases, NIAID, National Institutes of Health, Rockville, Maryland 20852, USA. garman@alpha.niaid.nih.gov
Organizational Affiliation: