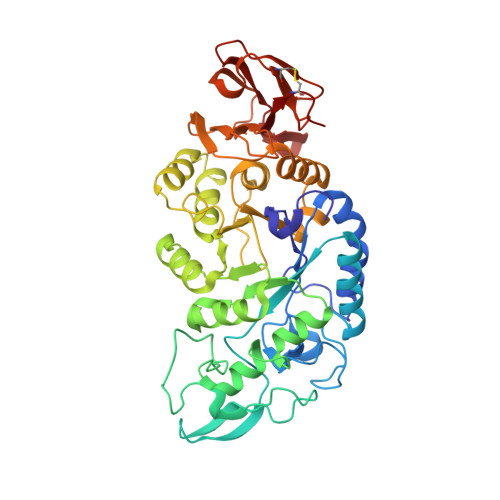
Differential Regulation of a Hyperthermophilic alpha-Amylase with a Novel (Ca,Zn) Two-metal Center by Zinc
Linden, A., Mayans, O., Meyer-Klaucke, W., Antranikian, G., Wilmanns, M.(2003) J Biological Chem 278: 9875-9884
- PubMed: 12482867
- DOI: https://doi.org/10.1074/jbc.M211339200
- Primary Citation of Related Structures:
1MWO, 1MXD, 1MXG - PubMed Abstract:
The crystal structure of the alpha-amylase from the hyperthermophilic archaeon Pyrococcus woesei was solved in the presence of three inhibitors: acarbose, Tris, and zinc. In the absence of exogenous metals, this alpha-amylase bound 1 and 4 molar eq of zinc and calcium, respectively. The structure reveals a novel, activating, two-metal (Ca,Zn)-binding site and a second inhibitory zinc-binding site that is found in the -1 sugar-binding pocket within the active site. The data resolve the apparent paradox between the zinc requirement for catalytic activity and its strong inhibitory effect when added in molar excess. They provide a rationale as to why this alpha-amylase, in contrast to commercially available alpha-amylases, does not require the addition of metal ions for full catalytic activity, suggesting it as an ideal target to maximize the efficiency of industrial processes like liquefaction of starch.
- European Molecular Biology Laboratory, Notkestrasse 85, D-22603 Hamburg, Germany.
Organizational Affiliation: