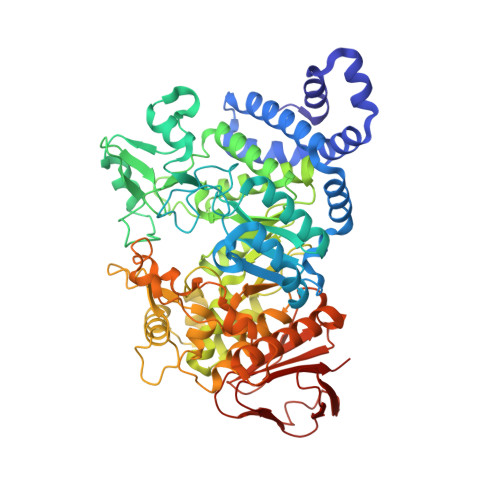
Oligosaccharide and Sucrose Complexes of Amylosucrase. STRUCTURAL IMPLICATIONS FOR THE POLYMERASE ACTIVITY
Skov, L.K., Mirza, O., Sprogoe, D., Dar, I., Remaud-Simeon, M., Albenne, C., Monsan, P., Gajhede, M.(2002) J Biological Chem 277: 47741-47747
- PubMed: 12364331 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M207860200
- Primary Citation Related Structures:
1MVY, 1MW0, 1MW1, 1MW2, 1MW3 - PubMed Abstract:
The glucosyltransferase amylosucrase is structurally quite similar to the hydrolase alpha-amylase. How this switch in functionality is achieved is an important and fundamental question. The inactive E328Q amylosucrase variant has been co-crystallized with maltoheptaose, and the structure was determined by x-ray crystallography to 2.2 A resolution, revealing a maltoheptaose binding site in the B'-domain somewhat distant from the active site. Additional soaking of these crystals with maltoheptaose resulted in replacement of Tris in the active site with maltoheptaose, allowing the mapping of the -1 to +5 binding subsites. Crystals of amylosucrase were soaked with sucrose at different concentrations. The structures at approximately 2.1 A resolution revealed three new binding sites of different affinity. The highest affinity binding site is close to the active site but is not in the previously identified substrate access channel. Allosteric regulation seems necessary to facilitate access from this binding site. The structures show the pivotal role of the B'-domain in the transferase reaction. Based on these observations, an extension of the hydrolase reaction mechanism valid for this enzyme can be proposed. In this mechanism, the glycogen-like polymer is bound in the widest access channel to the active site. The polymer binding introduces structural changes that allow sucrose to migrate from its binding site into the active site and displace the polymer.
- Protein Structure Group, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark. lsk@dfh.dk
Organizational Affiliation: