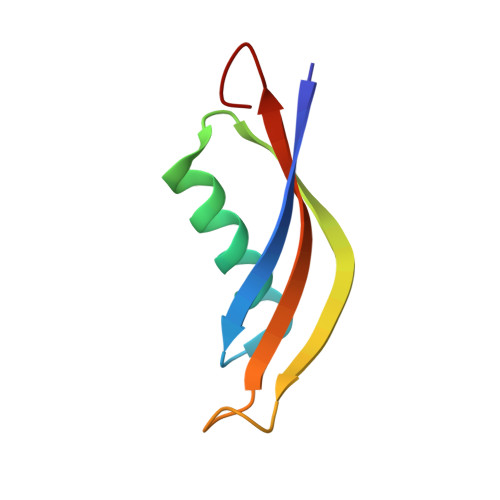
Crystal Structure of Halophilic Dodecin: A Novel, Dodecameric Flavin Binding Protein from Halobacterium salinarum
Bieger, B., Essen, L.-O., Oesterhelt, D.(2003) Structure 11: 375-385
- PubMed: 12679016
- DOI: https://doi.org/10.1016/s0969-2126(03)00048-0
- Primary Citation of Related Structures:
1MOG - PubMed Abstract:
A novel, 68 amino acid long flavoprotein called dodecin has been discovered in the proteome of Halobacterium salinarum by inverse structural genomics. The 1.7 A crystal structure of this protein shows a dodecameric, hollow sphere-like arrangement of the protein subunits. Unlike other known flavoproteins, which bind only monomeric flavin cofactors, the structure of the dodecin oligomer comprises six riboflavin dimers. The dimerization of these riboflavins along the re-faces is mediated by aromatic, antiparallel pi staggering of their isoalloxazine moieties. A unique aromatic tetrade is formed by further sandwiching of the riboflavin dimers between the indole groups of two symmetry-related Trp36s. So far, the dodecins represent the smallest known flavoproteins. Based on the structure and the wide spread occurrences in pathogenic and soil eubacteria, a function in flavin storage or protection against radical or oxygenic stress is suggested for the dodecins.
- Department of Membrane Biochemistry, Max Planck Institute for Biochemistry, Am Klopferspitz 18a, D-82152, Martinsried, Germany.
Organizational Affiliation: