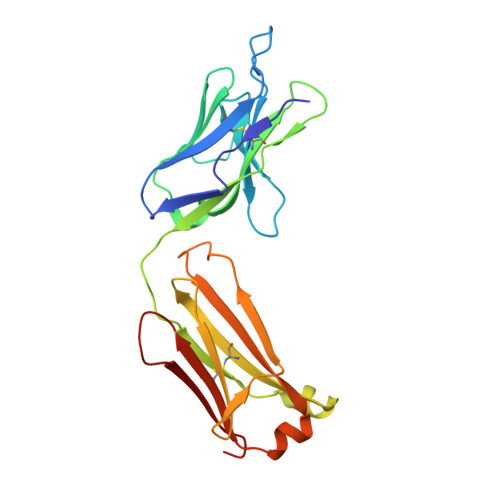
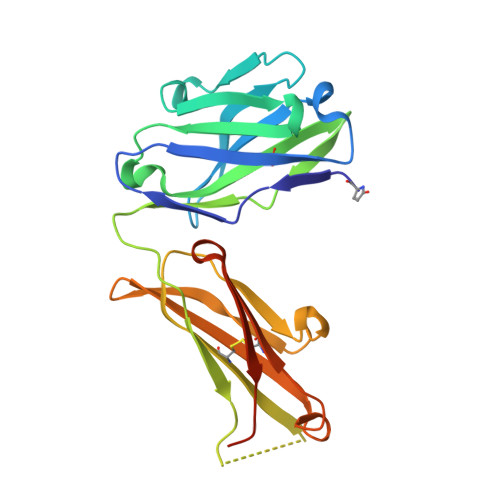
High-resolution crystal structure of the Fab-fragments of a family of mouse catalytic antibodies with esterase activity
Ruzheinikov, S.N., Muranova, T.A., Sedelnikova, S.E., Partridge, L.J., Blackburn, G.M., Murray, I.A., Kakinuma, H., Takashi, N., Shimazaki, K., Sun, J., Nishi, Y., Rice, D.W.(2003) J Mol Biology 332: 423-435
- PubMed: 12948492 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00902-1
- Primary Citation Related Structures:
1MH5, 1MIE, 1MJ7, 1MJ8, 1MJJ, 1MJU - PubMed Abstract:
The crystal structures of four related Fab fragments of a family of catalytic antibodies displaying differential levels of esterase activity have been solved in the presence and in the absence of the transition-state analogue (TSA) that was used to elicit the immune response. The electron density maps show that the TSA conformation is essentially identical, with limited changes on hapten binding. Interactions with the TSA explain the specificity for the D rather than the L-isomer of the substrate. Differences in the residues in the hapten-binding pocket, which increase hydrophobicity, appear to correlate with an increase in the affinity of the antibodies for their substrate. Analysis of the structures at the active site reveals a network of conserved hydrogen bond contacts between the TSA and the antibodies, and points to a critical role of two conserved residues, HisL91 and LysH95, in catalysis. However, these two key residues are set into very different contexts in their respective structures, with an apparent direct correlation between the catalytic power of the antibodies and the complexity of their interactions with the rest of the protein. This suggests that the catalytic efficiency may be controlled by contacts arising from a second sphere of residues at the periphery of the active site.
- Krebs Institute for Biomolecular Research, University of Sheffield, Firth Court, Western Bank, S10 2TN Sheffield, UK.
Organizational Affiliation: