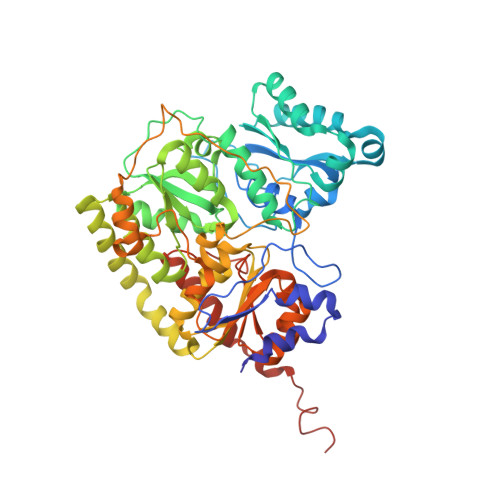
X-ray crystal structure of the nitrogenase molybdenum-iron protein from Clostridium pasteurianum at 3.0-A resolution.
Kim, J., Woo, D., Rees, D.C.(1993) Biochemistry 32: 7104-7115
- PubMed: 8393705 Search on PubMed
- DOI: https://doi.org/10.1021/bi00079a006
- Primary Citation Related Structures:
1MIO - PubMed Abstract:
The crystal structure of the nitrogenase molybdenum-iron (MoFe) protein from Clostridium pasteurianum (Cp1) has been determined at 3.0-A resolution by a combination of isomorphous replacement, molecular replacement, and noncrystallographic symmetry averaging. The structure of Cp1, including the two types of metal centers associated with the protein (the FeMo-cofactor and the P-cluster pair), is similar to that previously described for the MoFe-protein from Azotobacter vinelandii (Av1). Unique features of the Cp1 structure arise from the presence of an approximately 50-residue insertion in the alpha subunit and an approximately 50-residue deletion in the beta subunit. As a consequence, the FeMo-cofactor is more buried in Cp1 than in Av1, since the insertion is located on the surface above the FeMo-cofactor. The location of this insertion near the putative nitrogenase iron protein binding site provides a structural basis for the observation that the nitrogenase proteins from C. pasteurianum have low activity with complementary nitrogenase proteins isolated from other organisms. Mechanistic implications of the Cp1 structure for substrate entry/product release, substrate binding to the FeMo-cofactor, and electron- and proton-transfer reactions of nitrogenase are discussed.
- Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena 91125.
Organizational Affiliation: