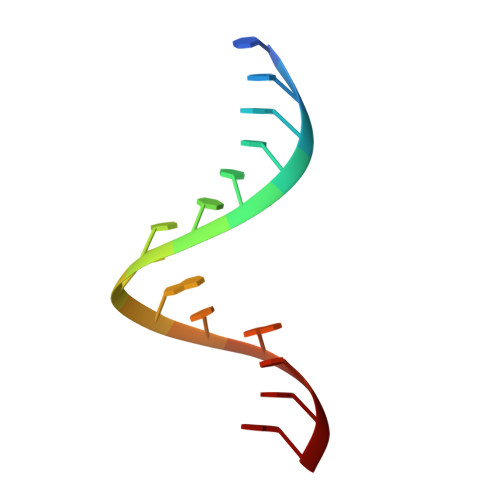
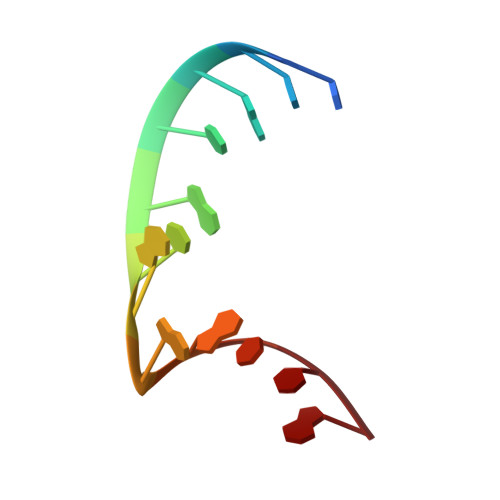
The crystal structure of a 26-nucleotide RNA containing a hook-turn
Szep, S., Wang, J., Moore, P.B.(2003) RNA 9: 44-51
- PubMed: 12554875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.2107303
- Primary Citation Related Structures:
1MHK - PubMed Abstract:
A crystal structure has been obtained for a 26-nucleotide RNA that contains the loop E sequence from Chromatium minutissimum. Rather than having a loop E-like conformation, it consists of an A-form helix that splits into two separate strands following a sheared A-G base pair. The backbone of the strand containing the G of the A-G pair makes a turn of almost 180 degrees in the space of two nucleotides, and then interacts with the minor groove of the helix from which it originates. Similar structures, which we call hook-turns, occur in 16S and 23S rRNAs. They are found at places where the two strands of a helix separate at an A/G juxtaposition to interact with other sequences.
- Department of Chemistry, Yale University, New Haven, Connecticut 06520, USA.
Organizational Affiliation: