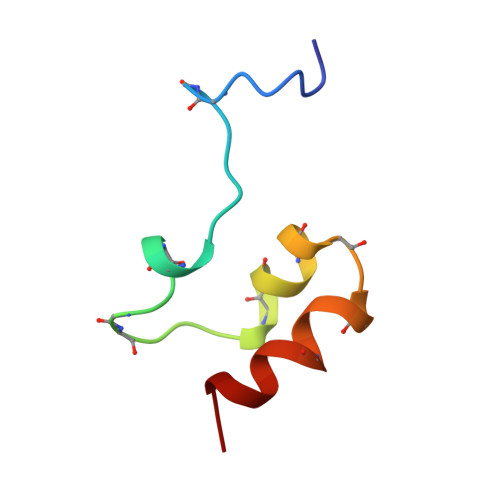
Identification of the phospholipid binding site in the vitamin K-dependent blood coagulation protein factor IX.
Freedman, S.J., Blostein, M.D., Baleja, J.D., Jacobs, M., Furie, B.C., Furie, B.(1996) J Biological Chem 271: 16227-16236
- PubMed: 8663165 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.271.27.16227
- Primary Citation Related Structures:
1MGX - PubMed Abstract:
The blood coagulation and regulatory proteins that contain gamma-carboxyglutamic acid are a part of a unique class of membrane binding proteins that require calcium for their interaction with cell membranes. Following protein biosynthesis, glutamic acids on these proteins are converted to gamma-carboxyglutamic acid (Gla) in a reaction that requires vitamin K as a cofactor. The vitamin K-dependent proteins undergo a conformational transition upon metal ion binding, but only calcium ions mediate protein-phospholipid interaction. To identify the site on Factor IX that is required for phospholipid binding, we have determined the three-dimensional structure of the Factor IX Gla domain bound to magnesium ions by NMR spectroscopy. By comparison of this structure to that of the Gla domain bound to calcium ions, we localize the membrane binding site to a highly ordered structure including residues 1-11 of the Gla domain. In the presence of Ca2+, Factor IX Gla domain peptides that contain the photoactivatable amino acid p-benzoyl-L-phenylalanine at positions 6 or 9 cross-link to phospholipid following irradiation, while peptides lacking this amino acid analog or with this analog at position 46 did not cross-link. These results indicate that the NH2 terminus of the Gla domain, specifically including leucine 6 and phenylalanine 9 in the hydrophobic patch, is the contact surface on Factor IX that interacts with the phospholipid bilayer.
- Center for Hemostasis and Thrombosis Research, Division of Hematology-Oncology, New England Medical Center, Boston, Massachusetts 02111, USA.
Organizational Affiliation: