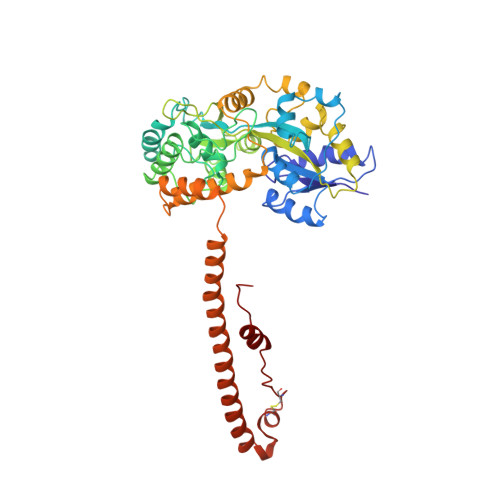
Crystal structure of human T cell leukemia virus type 1 gp21 ectodomain crystallized as a maltose-binding protein chimera reveals structural evolution of retroviral transmembrane proteins.
Kobe, B., Center, R.J., Kemp, B.E., Poumbourios, P.(1999) Proc Natl Acad Sci U S A 96: 4319-4324
- PubMed: 10200260 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.8.4319
- Primary Citation Related Structures:
1MG1 - PubMed Abstract:
Retroviral entry into cells depends on envelope glycoproteins, whereby receptor binding to the surface-exposed subunit triggers membrane fusion by the transmembrane protein (TM) subunit. We determined the crystal structure at 2.5-A resolution of the ectodomain of gp21, the TM from human T cell leukemia virus type 1. The gp21 fragment was crystallized as a maltose-binding protein chimera, and the maltose-binding protein domain was used to solve the initial phases by the method of molecular replacement. The structure of gp21 comprises an N-terminal trimeric coiled coil, an adjacent disulfide-bonded loop that stabilizes a chain reversal, and a C-terminal sequence structurally distinct from HIV type 1/simian immunodeficiency virus gp41 that packs against the coil in an extended antiparallel fashion. Comparison of the gp21 structure with the structures of other retroviral TMs contrasts the conserved nature of the coiled coil-forming region and adjacent disulfide-bonded loop with the variable nature of the C-terminal ectodomain segment. The structure points to these features having evolved to enable the dual roles of retroviral TMs: conserved fusion function and an ability to anchor diverse surface-exposed subunit structures to the virion envelope and infected cell surface. The structure of gp21 implies that the N-terminal fusion peptide is in close proximity to the C-terminal transmembrane domain and likely represents a postfusion conformation.
- St. Vincent's Institute of Medical Research, 41 Victoria Parade, Fitzroy, Victoria 3065, Australia.
Organizational Affiliation: