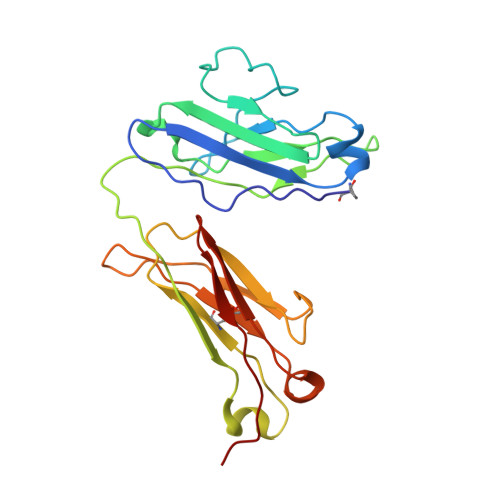
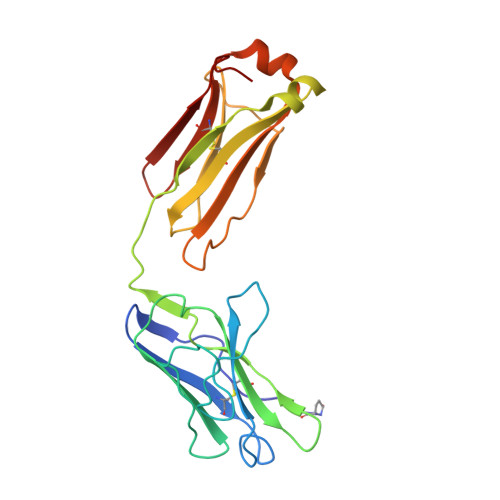
Three-dimensional structure of a hybrid light chain dimer: protein engineering of a binding cavity.
Ely, K.R., Herron, J.N., Edmundson, A.B.(1990) Mol Immunol 27: 101-114
- PubMed: 2108322 Search on PubMed
- DOI: https://doi.org/10.1016/0161-5890(90)90105-9
- Primary Citation Related Structures:
1MCW - PubMed Abstract:
An attempt was made to engineer a binding site and check its structure by X-ray analysis. Two human light chains (Mcg and Weir), with "variable" domain sequences differing in 36 positions, were hybridized into a heterologous dimer and crystallized in ammonium sulfate by the same procedure used for the trigonal form of the Mcg dimer. The three-dimensional structure of the hybrid was determined at 3.5-A resolution by difference Fourier analysis, interactive model building with computer graphics and crystallographic refinement. In the heterologous dimer, the Weir protein behaved as the structural analog of the heavy chain in an antigen binding fragment, while the Mcg protein assumed the role of the light chain component. The hybrid and the Mcg dimer were closely similar in overall structure, an observation probably correlated with the deliberate cleavage of the intrachain disulfide bond in the variable domain of the Weir protein during the hybridization procedure. Examination of the crystal structure of the hybrid suggested that the cleavage resulted in the relaxation of restraints which might otherwise have interfered with the formation of an Mcg-like dimer. There were six substitutions among the residues lining the binding cavities of the hybrid and Mcg dimer. These substitutions significantly affected the sizes, shapes and binding properties of the two cavities.
- Department of Biology, University of Utah, Salt Lake City 84112.
Organizational Affiliation: