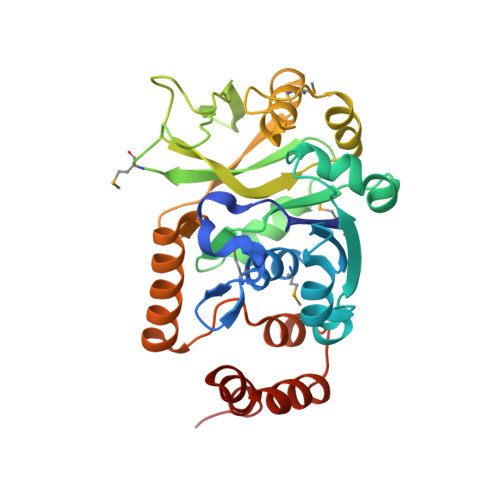
Crystal Structure of Escherichia coli Glucose-1-Phosphate Thymidylyltransferase (RffH) Complexed with dTTP and Mg2+
Sivaraman, J., Sauve, V., Matte, A., Cygler, M.(2002) J Biological Chem 277: 44214-44219
- PubMed: 12171937
- DOI: https://doi.org/10.1074/jbc.M206932200
- Primary Citation Related Structures:
1MC3 - PubMed Abstract:
The enzyme glucose-1-phosphate thymidylyltransferase (RffH), the product of the rffh gene, catalyzes one of the steps in the synthesis of enterobacterial common antigen (ECA), a cell surface glycolipid found in Gram-negative enteric bacteria. In Escherichia coli two gene products, RffH and RmlA, catalyze the same enzymatic reaction and are homologous in sequence; however, they are part of different operons and function in different pathways. We report the crystal structure of RffH bound to deoxythymidine triphosphate (dTTP), the phosphate donor, and Mg(2+), refined at 2.6 A to an R-factor of 22.3% (R(free) = 28.4%). The crystal structure of RffH shows a tetrameric enzyme best described as a dimer of dimers. Each monomer has an overall alpha/beta fold and consists of two domains, a larger nucleotide binding domain (residues 1-115, 222-291) and a smaller sugar-binding domain (116-221), with the active site located at the domain interface. The Mg(2+) ion is coordinated by two conserved aspartates and the alpha-phosphate of deoxythymidine triphosphate. Its location corresponds well to that in a structurally similar domain of N-acetylglucosamine-1-phosphate uridylyltransferase (GlmU). Analysis of the RffH, RmlA, and GlmU complexes with substrates and products provides an explanation for their different affinities for Mg(2+) and leads to a proposal for the dynamics along the reaction pathway.
- Department of Biochemistry, McGill University, Montréal, Québec H3G 1Y6, Canada.
Organizational Affiliation: