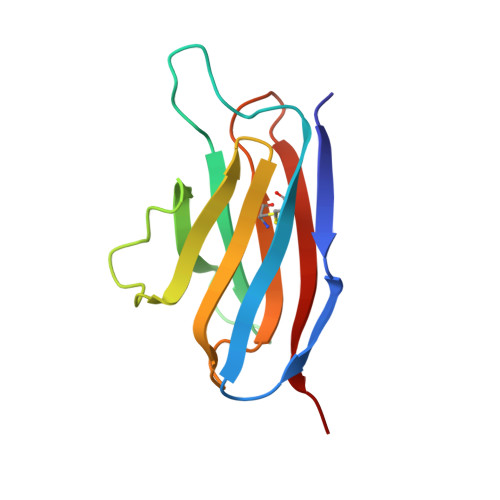
Solution structure of an isolated antibody VL domain.
Constantine, K.L., Friedrichs, M.S., Metzler, W.J., Wittekind, M., Hensley, P., Mueller, L.(1994) J Mol Biology 236: 310-327
- PubMed: 8107112
- DOI: https://doi.org/10.1006/jmbi.1994.1137
- Primary Citation of Related Structures:
1MAJ, 1MAK - PubMed Abstract:
The solution structure of the isolated VL domain of the anti-digoxin antibody 26-10 has been determined using data derived from heteronuclear multi-dimensional nuclear magnetic resonance (n.m.r.) experiments. Analytical ultracentrifugation and n.m.r. data demonstrate that the VL domain is only weakly associating (Kd = 2.5 (+/- 0.7) mM) and that it experiences a rapid monomer/dimer equilibrium under the n.m.r. experimental conditions. Therefore, the results reported here represent the first structure determination of an antibody VL domain in the absence of fixed quaternary interactions. The structure determination is based on 930 proton-proton distance constraints, 113 dihedral angle constraints, and 46 hydrogen bond constraints. Eighty initial structures were calculated with the variable target function program DIANA; of these, 31 were accepted on the basis of satisfaction of constraints (no distance constraint violations > 0.5 A; target function < 3.0 A2). Accepted DIANA structures were refined by restrained energy minimization using the X-PLOR program. The 15 best energy-minimized DIANA structures were chosen as a representative ensemble of solution conformations. The average root-mean-square differences (r.m.s.d.) between the individual structures of this ensemble and the mean coordinates is 0.85 (+/- 0.10) A for all backbone atoms and 1.29 (+/- 0.10) A for all heavy atoms. For beta-strands A, B, C, D, E and F, the average backbone atom r.m.s.d. to the mean structure is 0.46 (+/- 0.06) A. A higher-resolution ensemble, with all backbone atom and all heavy atom r.m.s.d.s. to the mean coordinates of 0.54 (+/- 0.08) A and 0.98 (+/- 0.12) A, respectively, was obtained by X-PLOR simulated annealing refinement of the 15 energy-minimized DIANA structures. A detailed analysis of the original ensemble of 15 energy-minimized DIANA structures is presented, as this ensemble retains a broader, and possibly more realistic, sampling of conformation space. The backbone atom and all heavy atom r.m.s.d.s between the mean energy-minimized DIANA structure and the X-ray derived coordinates of the VL domain within the Fab/digoxin complex are 1.05 A and 1.56 A, respectively. Subtle differences between the solution and X-ray structures occur primarily in CDR2, CDR3, beta-strands A, F and G, and localized regions of hydrophobic packing. Overall, these results demonstrate that the 26-10 VL domain conformation is determined primarily by intradomain interactions, and that quaternary VL-VH association induces relatively minor conformational adjustments.
- Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, NJ 08543.
Organizational Affiliation: