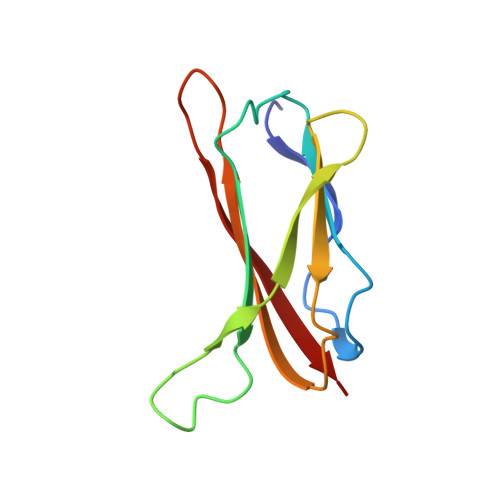
Solution structure of CopC: a cupredoxin-like protein involved in copper homeostasis
Arnesano, F., Banci, L., Bertini, I., Thompsett, A.R.(2002) Structure 10: 1337-1347
- PubMed: 12377120 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00858-4
- Primary Citation Related Structures:
1M42 - PubMed Abstract:
The structure of the metal-free form of CopC, a protein involved in copper homeostasis, has been obtained. The fold is a Greek key beta barrel similar to that of functionally unrelated blue copper proteins but with important structural variations. The protein binds one equivalent of copper (II) with relatively high affinity and contains a cluster of conserved residues (His1, Glu27, Asp89, and His91) which could form a water-accessible metal binding site. The structure also reveals a loop containing the M(X)(n)M motif which is present in a number of proteins also involved in copper homeostasis. The present structure represents a link between copper-trafficking proteins and cupredoxins. Within a structural and genomic analysis, the role of CopC in copper trafficking is discussed.
- Magnetic Resonance Center (CERM) and Department of Chemistry, University of Florence, Sesto Fiorentino, Florence, Italy.
Organizational Affiliation: