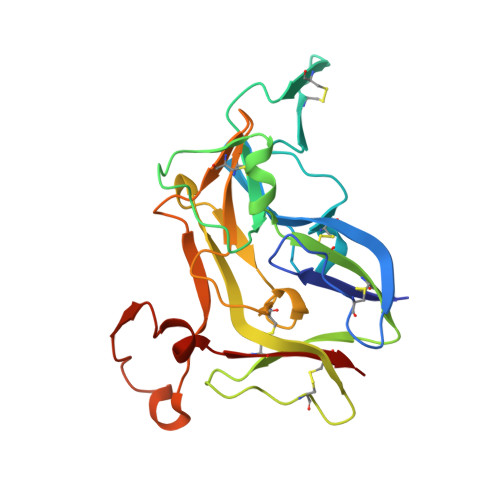
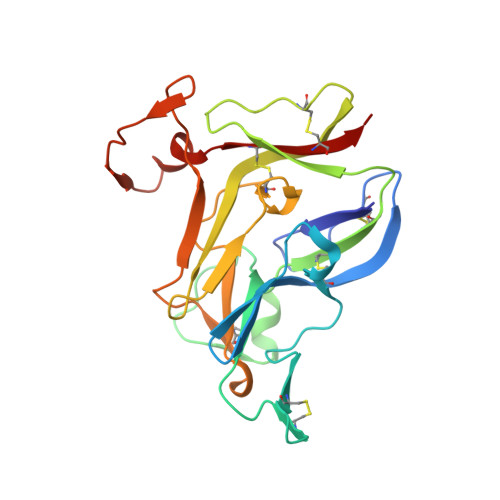
Crystal Structure of NC1 Domains. Structural Basis for Type IV Collagen Assembly in Basement Membranes
Sundaramoorthy, M., Meiyappan, M., Todd, P., Hudson, B.G.(2002) J Biological Chem 277: 31142-31153
- PubMed: 11970952
- DOI: https://doi.org/10.1074/jbc.M201740200
- Primary Citation of Related Structures:
1M3D - PubMed Abstract:
Type IV collagen, which is present in all metazoan, exists as a family of six homologous alpha(IV) chains, alpha1-alpha6, in mammals. The six chains assemble into three different triple helical protomers and self-associate as three distinct networks. The network underlies all epithelia as a component of basement membranes, which play important roles in cell adhesion, growth, differentiation, tissue repair and molecular ultrafiltration. The specificity of both protomer and network assembly is governed by amino acid sequences of the C-terminal noncollagenous (NC1) domain of each chain. In this study, the structural basis for protomer and network assembly was investigated by determining the crystal structure of the ubiquitous [(alpha1)(2).alpha2](2) NC1 hexamer of bovine lens capsule basement membrane at 2.0 A resolution. The NC1 monomer folds into a novel tertiary structure. The (alpha1)(2).alpha2 trimer is organized through the unique three-dimensional domain swapping interactions. The differences in the primary sequences of the hypervariable region manifest in different secondary structures, which determine the chain specificity at the monomer-monomer interfaces. The trimer-trimer interface is stabilized by the extensive hydrophobic and hydrophilic interactions without a need for disulfide cross-linking.
- Department of Biochemistry and Molecular Biology, University of Kansas Medical Center, Kansas City, KS 66160-7421, USA.
Organizational Affiliation: