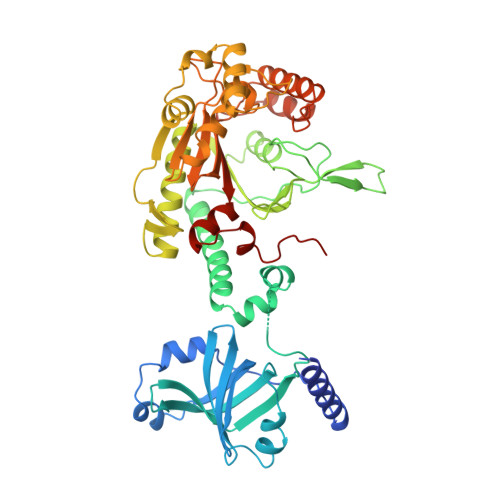
The crystal structure of the lysyl-tRNA synthetase (LysU) from Escherichia coli
Onesti, S., Miller, A.D., Brick, P.(1995) Structure 3: 163-176
- PubMed: 7735833 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00147-2
- Primary Citation Related Structures:
1LYL - PubMed Abstract:
Lysyl-tRNA synthetase catalyzes the attachment of the amino acid lysine to the cognate tRNA. The enzyme is a member of the class II amino-acyl-tRNA synthetases; the crystal structures of the seryl- and aspartyl-tRNA synthetases from this class are already known. Lysyl-tRNA synthetase shows extensive sequence homology with aspartyl-tRNA synthetase. In Escherichia coli there are two isoforms of the enzyme, LysS and LysU. Unlike LysS, which is synthesized under normal growth conditions, LysU is the product of a normally silent gene which is overexpressed under extreme physiological conditions (such as heat-shock), and can synthesize a number of adenyl dinucleotides (in particular AppppA). These dinucleotides have been proposed to act as modulators of the heat-shock response and stress response. The crystal structure of E. coli LysU has been determined to 2.8 A resolution, with lysine bound to the active site. The protein is a homodimer, with a rather extended dimer interface spanning the entire length of the molecule. Each monomer consists of two domains: a smaller N-terminal domain which binds the tRNA anticodon, and a larger C-terminal domain with the topology characteristic of the catalytic domain found in class II synthetases. A comparison of the LysU crystal structure with the structures of seryl- and aspartyl-tRNA synthetases enables a conserved core to be identified. The structural homology with the aspartyl-tRNA synthetase extends to include the anticodon-binding domain. When the active sites of lysyl-, aspartyl- and seryl-tRNA synthetases are compared, a number of catalytically important residues are conserved and a similar extended network of hydrogen bonds can be observed in the amino acid binding pocket in all three structures, although the details may differ. The lysine substrate is involved in an extended network of hydrogen bonds and polar interactions, with the side chain amino group forming a salt bridge with Glu428. The binding of ATP to LysU can be modelled on the basis of the aspartyl-tRNA synthetase-ATP complex, but the tRNA acceptor stem interaction for LysU cannot be easily modelled by similar extrapolation.
- Blackett Laboratory, Imperial College, London, UK.
Organizational Affiliation: