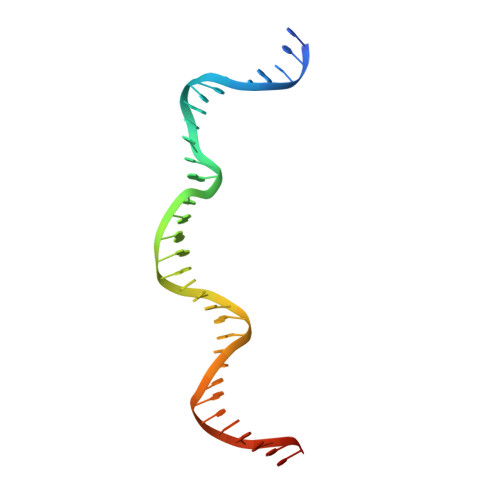
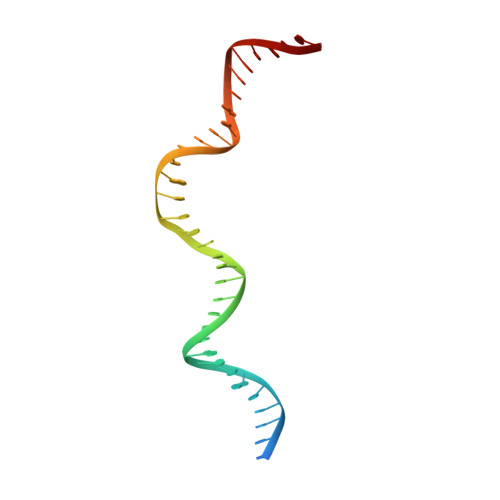
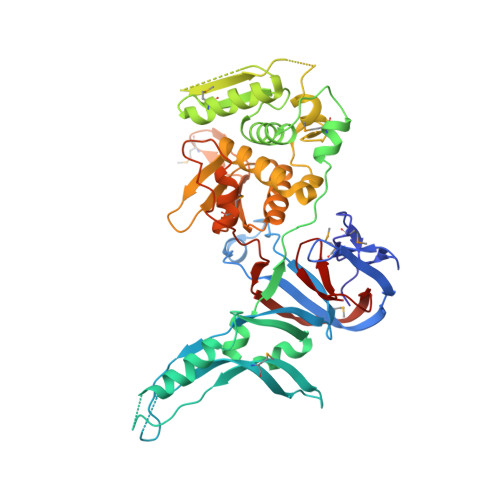
Crystal structure of the intein homing endonuclease PI-SceI bound to its recognition sequence.
Moure, C.M., Gimble, F.S., Quiocho, F.A.(2002) Nat Struct Biol 9: 764-770
- PubMed: 12219083 Search on PubMed
- DOI: https://doi.org/10.1038/nsb840
- Primary Citation Related Structures:
1LWS, 1LWT - PubMed Abstract:
The first X-ray structures of an intein-DNA complex, that of the two-domain homing endonuclease PI-SceI bound to its 36-base pair DNA substrate, have been determined in the presence and absence of Ca(2+). The DNA shows an asymmetric bending pattern, with a major 50 degree bend in the endonuclease domain and a minor 22 degree bend in the splicing domain region. Distortions of the DNA bound to the endonuclease domain cause the insertion of the two cleavage sites in the catalytic center. DNA binding induces changes in the protein conformation. The two overlapping non-identical active sites in the endonucleolytic center contain two Ca(+2) ions that coordinate to the catalytic Asp residues. Structure analysis indicates that the top strand may be cleaved first.
- Howard Hughes Medical Institute, Baylor College of Medicine, Houston, Texas 77030, USA.
Organizational Affiliation: