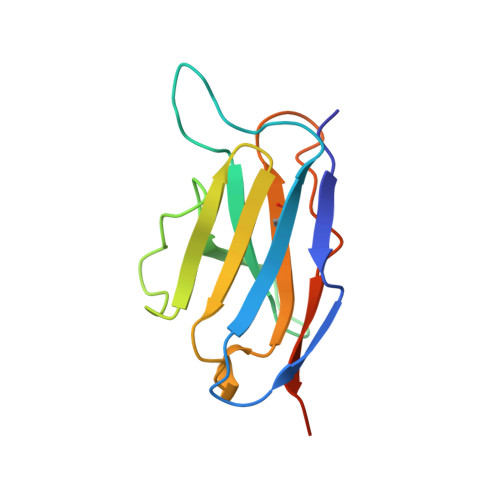
Variable domain structure of kappaIV human light chain Len: high homology to the murine light chain McPC603.
Huang, D.B., Chang, C.H., Ainsworth, C., Johnson, G., Solomon, A., Stevens, F.J., Schiffer, M.(1997) Mol Immunol 34: 1291-1301
- PubMed: 9683271
- DOI: https://doi.org/10.1016/s0161-5890(98)00002-9
- Primary Citation of Related Structures:
1LVE - PubMed Abstract:
Antibody light chains of the kappa subgroup are the predominant light chain component in human immune responses and are used almost exclusively in the antibody repertoire of mice. Human kappa light chains comprise four subgroups. To date, all crystallographic studies of human kappa light chains were carried out on proteins of the kappaI subgroup. The light chain produced by multiple myeloma patient Len. was of the kappaIV subgroup, it differed by only one residue from the germ-line gene encoded protein. The variable domain fragment of the light chain was crystallized from ammonium sulfate in space group C222(1). The crystal structure was determined by molecular replacement and refined at 1.95 A resolution to an R-factor of 0.15. Protein Len has six additional residues in its CDR1 segment compared to the kappaI proteins previously characterized. The kappaIV variable domain, Len, differs in only 23 of 113 residues from murine kappa light chain McPC603. The RMS deviation upon superimposing their alpha-carbons was 0.69 A. The CDR1 segment of the human and murine variable domains have the same length and conformation although their amino acid sequences differ in 5 out of 17 residues. Structural features were identified that could account for the significantly higher stability of the human kappaIV protein relative to its murine counterpart. This human kappaIV light chain structure is the closest human homolog to a murine light chain and can be expected to facilitate detailed structural comparisons necessary for effective humanization of murine antibodies.
- Center for Mechanistic Biology and Biotechnology, Argonne National Laboratory, IL 60439, USA.
Organizational Affiliation: