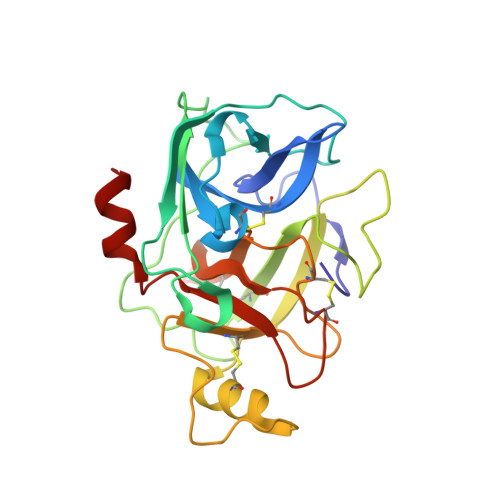
The Crystal Structure of Human alpha1-Tryptase Reveals a Blocked Substrate-binding Region
Marquardt, U., Zettl, F., Huber, R., Bode, W., Sommerhoff, C.P.(2002) J Mol Biology 321: 491-502
- PubMed: 12162961 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00625-3
- Primary Citation Related Structures:
1LTO - PubMed Abstract:
Human mast cell tryptases represent a subfamily of trypsin-like serine proteinases implicated in asthma. Unlike beta-tryptases, alpha-tryptases apparently are proteolytically inactive. We have solved the 2.2A crystal structure of mature human alpha1-tryptase. It reveals a frame-like tetrameric architecture that, surprisingly, does not require heparin-binding for stability. In marked contrast to beta2-tryptase, the Ser214-Gly219 segment, which normally provides the template for substrate binding, is kinked in alpha-tryptase, thereby blocking its non-primed subsites. This so far unobserved subsite distortion is incompatible with productive substrate binding and processing. alpha-Tryptase apparently is trapped in this off-conformation by repulsions and attractions of the Asp216 side-chain. However, proteolytic activity could be generated by an induced-fit mechanism.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried bei, München, Germany.
Organizational Affiliation: