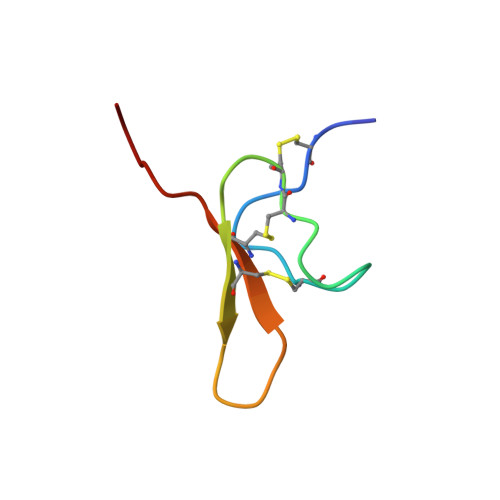
Recombinant production and solution structure of PcTx1, the specific peptide inhibitor of ASIC1a proton-gated cation channels
Escoubas, P., Bernard, C., Lambeau, G., Lazdunski, M., Darbon, H.(2003) Protein Sci 12: 1332-1343
- PubMed: 12824480 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0307003
- Primary Citation Related Structures:
1LMM - PubMed Abstract:
Acid-sensing ion channels (ASICs) are thought to be important ion channels, particularly for the perception of pain. Some of them may also contribute to synaptic plasticity, learning, and memory. Psalmotoxin 1 (PcTx1), the first potent and specific blocker of the ASIC1a proton-sensing channel, has been successfully expressed in the Drosophila melanogaster S2 cell recombinant expression system used here for the first time to produce a spider toxin. The recombinant toxin was identical in all respects to the native peptide, and its three-dimensional structure in solution was determined by means of (1)H 2D NMR spectroscopy. Surface characteristics of PcTx1 provide insights on key structural elements involved in the binding of PcTx1 to ASIC1a channels. They appear to be localized in the beta-sheet and the beta-turn linking the strands, as indicated by electrostatic anisotropy calculations, surface charge distribution, and the presence of residues known to be implicated in channel recognition by other inhibitor cystine knot (ICK) toxins.
- Institut de Pharmacologie Moléculaire et Cellulaire (IPMC), CNRS UMR 6097, Sophia-Antipolis, 06560 Valbonne, France.
Organizational Affiliation: